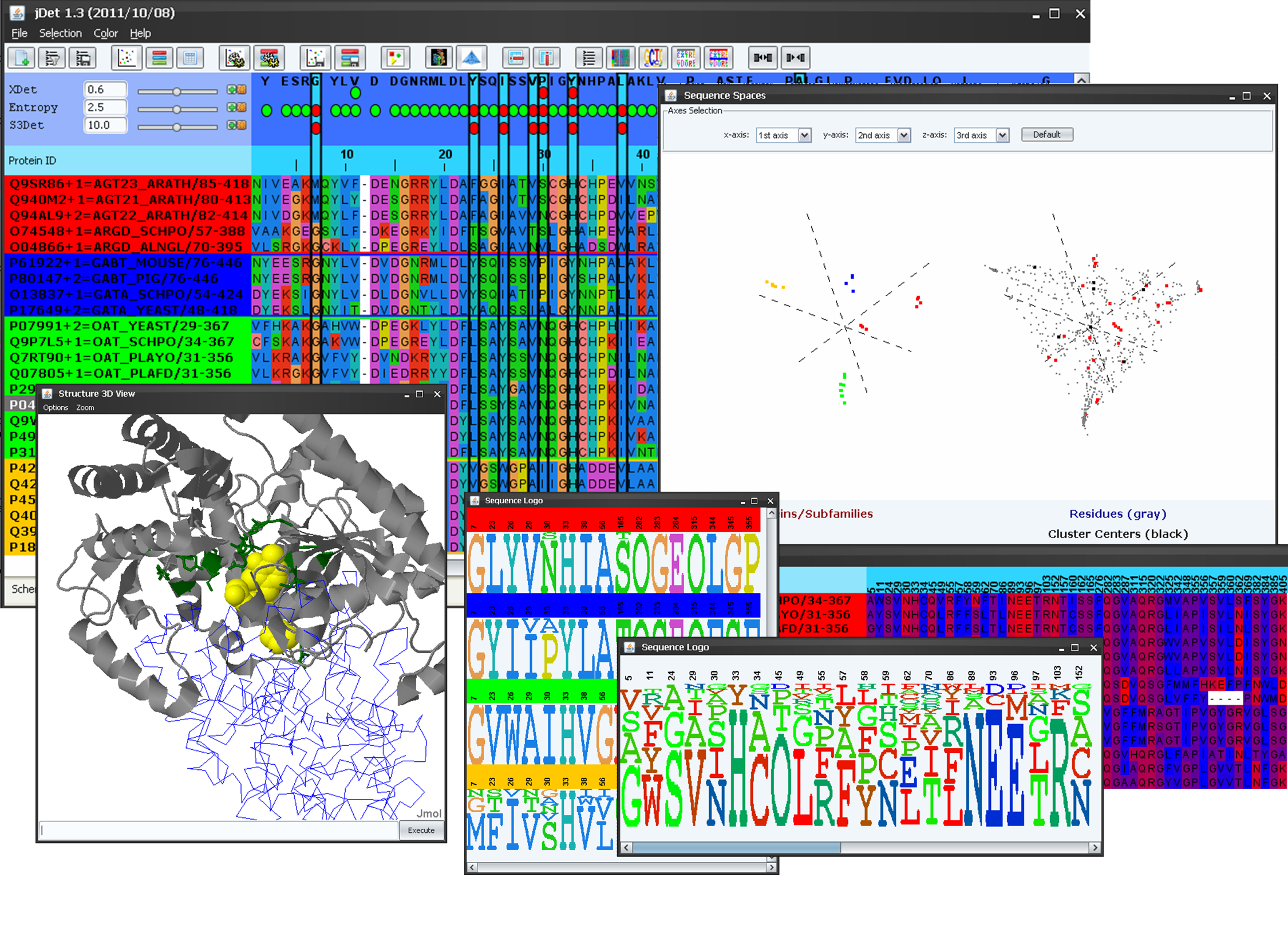
JDet is a multiplatform software for the interactive calculation and visualization of function-related conservation patterns in multiple sequence alignments and structures. It contains the set of tools and features we consider critical for the daily work with this kind of data, and that previously were disseminated in different packages and web servers. The package includes two of our recently developed programs for extracting this kind of information from protein alignments. JDet can be extended via plugins to implement other methods or parse their results.

For a general description of the package and the biological problem which prompted its development, as well as more information on multiple sequence alignments (MSAs), conserved positions, specificity determining positions (SDPs), etc., please take a look at the following article: Muth et al. (2012). Bioinformatics 28 (4): 584-586.; and the references cited there, specially Pazos et al. (2006) Bioinformatics. 22(12):1440-1448 and Rausell et al. (2010) PNAS. 107(5)1995-2000
Here you can find on-line versions of the user manual and the guided tutorial. These are also included in the dowloadable distribution.
 ).
).
Please cite the following reference when reporting any data or image generated with the program:
JDet includes binary distributions of two functional residue detection programs, Xdet and S3Det. These binaries can be also used outside the JDet environment, directly from the command line, e.g. for massive/batch runs (see Documentation above).
 ).
).
JDet uses Java3DTM, for research and instructional Use
JDet also includes the Jmol plugin:
Other third-party libraries are referenced in JDet's site at Google Code
For general questions/queries about the software, please contact Florencio Pazos ( ), for technical questions contact Juan A. Garcia Martin (
), for technical questions contact Juan A. Garcia Martin ( ).
).