BT_Metabolomics (442 tools)
| Name | Terms | Description | Date | #cit TOT | cit/ year | cit slope | cit pattern | cit rev |
|---|---|---|---|---|---|---|---|---|
| Metascape | Metabolomics Molecular interactions, pathways and networks Proteomics | Portal designed to provide a comprehensive gene list annotation and analysis resource for experimental biologists. | 2019-12 | 2965 | 741.2 | 406.9 | 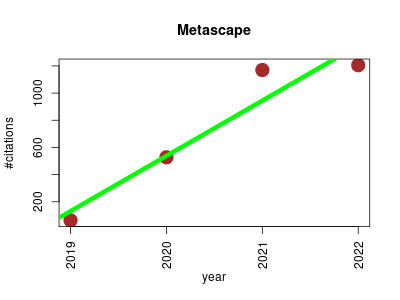 | 51 |
| MetaboAnalyst 4.0 | Metabolomics Metagenomics Small molecules | Statistical, functional and integrative analysis of metabolomics data , interpretation, and integration with other omics data. | 2018-07 | 1369 | 273.8 | 53.2 | 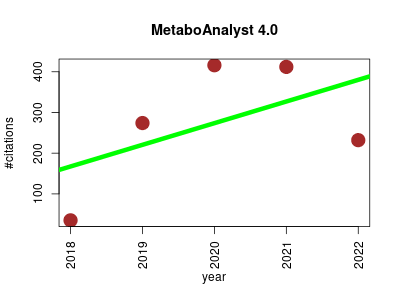 | 61 |
| HMDB | Metabolomics Model organisms NMR Small molecules Surgery | Freely available electronic database containing detailed information about small molecule metabolites found in the human body. The database supports extensive text, sequence, chemical structure and relational query searches. | 2005-12 | 1360 | 75.6 | 8.3 |  | 247 |
| proFIA | Metabolomics | Flow Injection Analysis coupled to High-Resolution Mass Spectrometry is a promising approach for high-throughput metabolomics. FIA-HRMS data, however, cannot be pre-processed with current software tools which rely on liquid chromatography separation, or handle low resolution data only. Here we present the package that implements a new methodology to pre-process FIA-HRMS raw data (netCDF, mzData, mzXML, and mzML) and generates the peak table. | 2015-01 | 1346 | 168.2 | 24.2 | 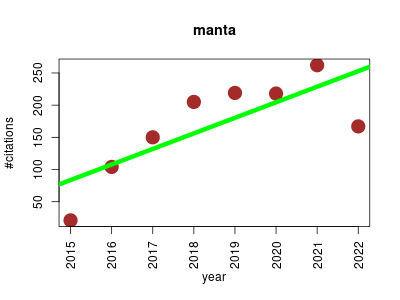 | 46 |
| flagme | Metabolomics | Fragment-level analysis of gas chromatography - mass spectrometry metabolomics data. | 2015-01 | 1346 | 168.2 | 24.2 |  | 46 |
| statTarget | Data quality management Metabolomics Proteomics experiment | An easy to use tool provide a graphical user interface for quality control based shift signal correction, integration of metabolomic data from multi-batch experiments, and the comprehensive statistic analysis in non-targeted or targeted metabolomics. | 2015-01 | 1346 | 168.2 | 24.2 |  | 46 |
| MetaboAnalyst 2.0 | MRI Metabolomics Molecular interactions, pathways and networks NMR Small molecules | Web-based pipeline for metabolomic data processing, statistical analysis and functional interpretation. It performs data processing and normalization for various metabolomic data types. It provides various univariate and multivariate statistical analysis for two/multi-group, one/two-factor, as well time-series data. | 2009-08 | 1130 | 80.7 | 9.1 | 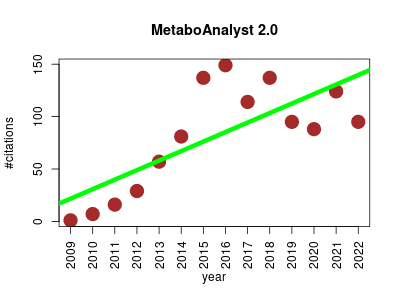 | 63 |
| GNPS | Biochemistry Biotechnology Lipids Metabolomics Proteomics experiment | GNPS is a web-based mass spectrometry ecosystem that aims to be an open-access knowledge base for community-wide organization and sharing of raw, processed, or annotated fragmentation mass spectrometry data (MS/MS). GNPS aids in identification and discovery throughout the entire life cycle of data; from initial data acquisition/analysis to post publication. | 2016-09 | 951 | 135.9 | 41.4 | 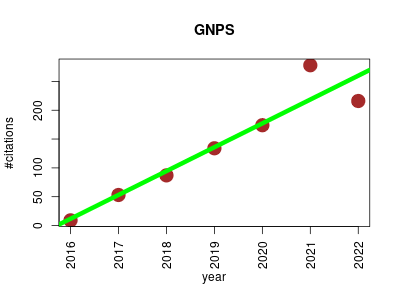 | 138 |
| mixomics | Data visualisation Machine learning Metabolomics Molecular interactions, pathways and networks Omics Proteomics Statistics and probability Transcriptomics | The tool offers a wide range of multivariate methods for the exploration and integration of biological datasets with a particular focus on variable selection. | 2009-01 | 923 | 65.9 | 18.3 | 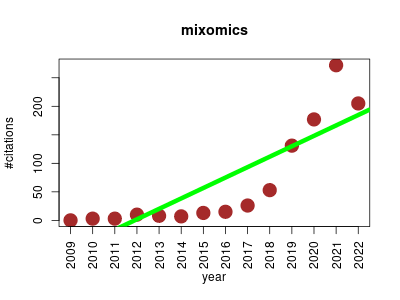 | 91 |
| Heatmapper | Data visualisation Metabolomics Proteomics Transcriptomics | Freely available web server that allows users to interactively visualize their data in the form of heat maps through an easy-to-use graphical interface. | 2016-07 | 680 | 97.1 | 33.8 | 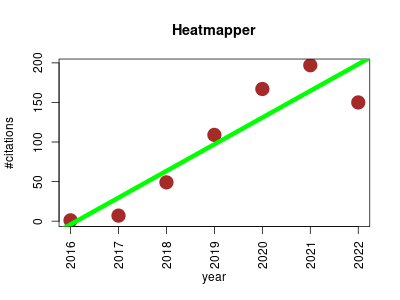 | 7 |
| MassBank | Metabolomics | MassBank is a public repository of mass spectra of small chemical compounds for life sciences (3000 Da). The database contains spectra from EI‐MS, fast atom bombardment MS, electrospray ionization (ESI)‐MSn data of thousands of authentic standards, EI‐MS and other‐MS data of thousands of volatile natural and synthetic compounds, and ESI‐MS2 data of synthetic drugs contributed by research groups worldwide. ESI‐MS2 data were analyzed under nonstandardized, independent experimental conditions. MassBank users can access either all of the MassBank data or a subset of the data by specifying one or more experimental conditions. In a spectral search to retrieve mass spectra similar to a query mass spectrum, the similarity score is calculated by a weighted cosine correlation in which weighting exponents on peak intensity and the mass‐to‐charge ratio are optimized to the ESI‐MS2 data. MassBank is useful for the identification of chemical compounds and the publication of experimental data. | 2010-01 | 573 | 44.1 | 5.9 | 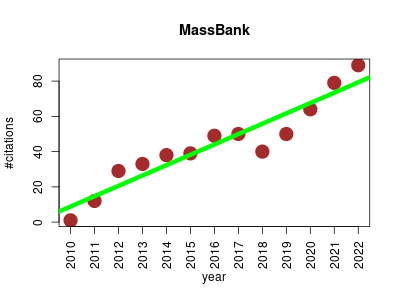 | 108 |
| CAMERA | Metabolomics | Annotation of peaklists generated by xcms, rule based annotation of isotopes and adducts, isotope validation, EIC correlation based tagging of unknown adducts and fragments. | 2012-01 | 358 | 32.5 | 5.1 | 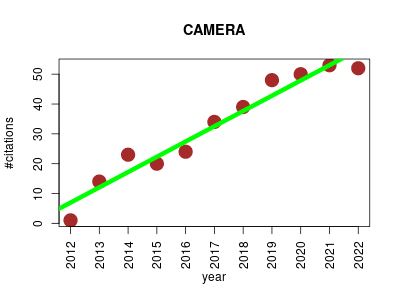 | 31 |
| rWikiPathways | Metabolomics Molecular interactions, pathways and networks Rare diseases | Use this package to interface with the WikiPathways API. | 2018-01 | 337 | 67.4 | 21.2 | 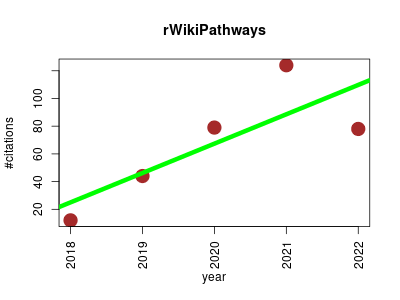 | 41 |
| COBRA Toolbox | Cheminformatics Metabolomics Molecular interactions, pathways and networks | Constraint-based reconstruction and analysis (COBRA) provides a molecular mechanistic framework for integrative analysis of experimental molecular systems biology data and quantitative prediction of physicochemically and biochemically feasible phenotypic states. | 2019-03 | 289 | 72.2 | 22.9 | 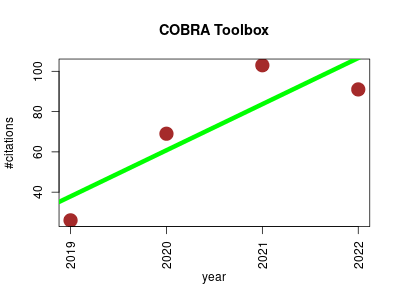 | 42 |
| MetaboLights | Metabolomics | Database for Metabolomics experiments and derived information. | 2013-01 | 257 | 25.7 | 1.1 | 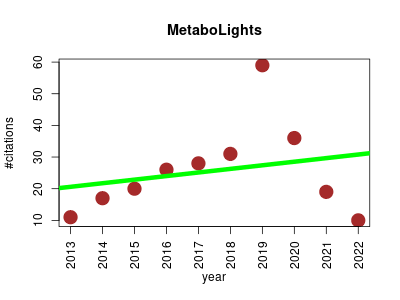 | 47 |
| MSEA | Genetics Metabolomics Molecular interactions, pathways and networks Small molecules | Performs enrichment analyses for (primarily human) metabolomic studies. It identifies patterns of metabolite concentration changes in a biologically meaningful context. It uses a library of ~6300 predefined metabolite sets from pathways, disease signatures, genetic traits, and cellular/tissue locations. It also facilitates conversion between metabolite common names, synonyms and other database identifies. | 2010-05 | 252 | 19.4 | 2.7 | 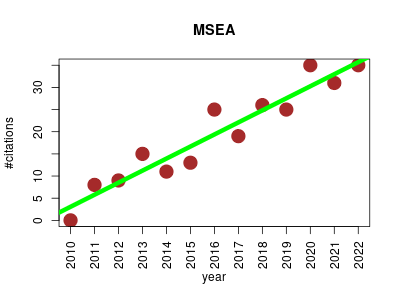 | 40 |
| FerrDb | Metabolomics Molecular biology Oncology Pathology Small molecules | FerrDb is a manually curated resource for regulators and markers of ferroptosis and ferroptosis-disease associations. | 2020-01 | 218 | 72.7 | 68.0 | 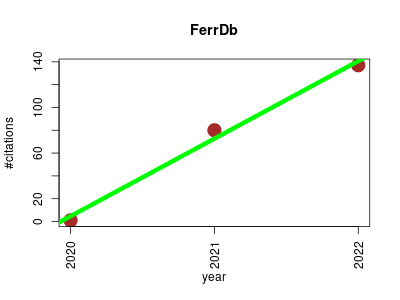 | 4 |
| MIBiG 2.0 | Drug discovery Metabolomics Metagenomics Microbial ecology Small molecules | A repository for biosynthetic gene clusters of known function. | 2020-01 | 181 | 60.3 | 19.0 | 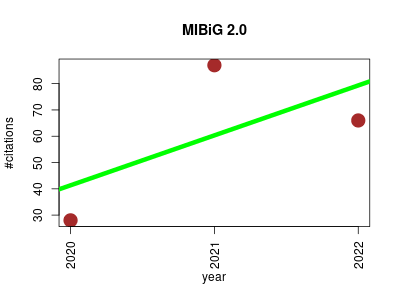 | 34 |
| CoNet app | Metabolomics Metagenomics Metatranscriptomics Microbial ecology Model organisms | inference of biological association networks using Cytoscape. | 2016-01 | 176 | 25.1 | 6.4 | 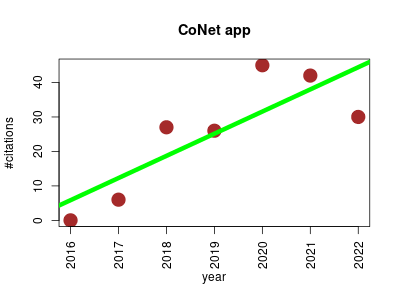 | 14 |
| MetaboliteDetector | Metabolomics Small molecules | QT4 based software package for the analysis of GC-MS based metabolomics data. The software is especially intended for the analysis of high resoluted GC-MS chromatograms which accumulate during high throughput based metabolmics experiments. | 2009-05 | 162 | 11.6 | 1.3 | 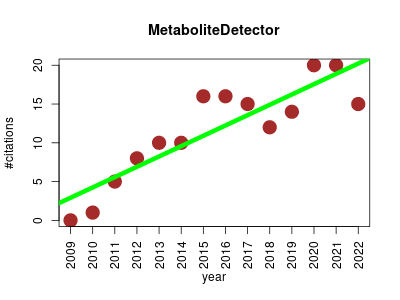 | 10 |
| Workflow4Metabolomics | Computational biology Metabolomics Systems biology | First fully open-source and collaborative online platform for computational metabolomics. It includes preprocessing, normalization, quality control, statistical analysis of LC/MS, FIA-MS, GC/MS and NMR data. | 2015-05 | 158 | 19.8 | 4.3 | 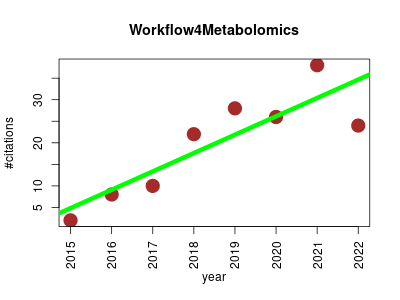 | 22 |
| BiG-SCAPE | Bioinformatics Mapping Metabolomics Microbial ecology Phylogeny | A computational framework to explore large-scale biosynthetic diversity. | 2020-01 | 158 | 52.7 | 5.5 | 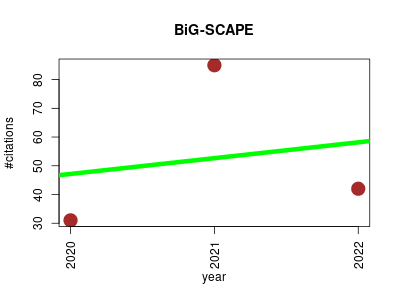 | 32 |
| OpenMS | Metabolomics Proteomics Proteomics experiment | Open source library and a collection of tools and interfaces for the analysis of mass spectrometry data. Includes over 200 standalone (TOPP) tools that can be combined to a workflow with the integrated workflow editor TOPPAS. Raw and intermediate mass spectrometry data can be visualised with the included viewer TOPPView. | 2011-01 | 157 | 13.1 | 6.1 | 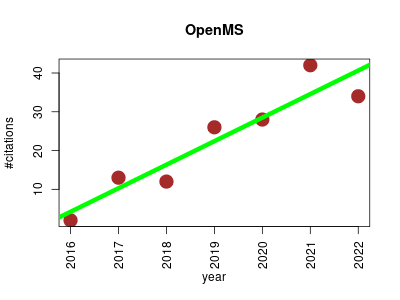 | 31 |
| MetaNetX | Biochemistry Cheminformatics Metabolomics Molecular interactions, pathways and networks Systems biology | MetaNetX/MNXref allows to access, analyse and manipulate genome-scale metabolic networks as well as biochemical pathways. It consistently integrates data from various public resources and makes the data accessible in a standardized format using a common namespace. Models can be interactively compared, analysed (e.g. detection of dead-end metabolites/reactions, flux balance analysis or simulation of reaction/gene knockouts), manipulated and exported. Users can upload their own metabolic models. | 2013-03 | 156 | 15.6 | 2.1 |  | 20 |
| CFM-ID | Machine learning Metabolomics Small molecules Structure prediction | CFM-ID (Competitive Fragmentation Modeling) is a web server for annotation, spectrum prediction and metabolite identification from tandem mass spectra data. | 2014-07 | 150 | 16.7 | 3.0 | 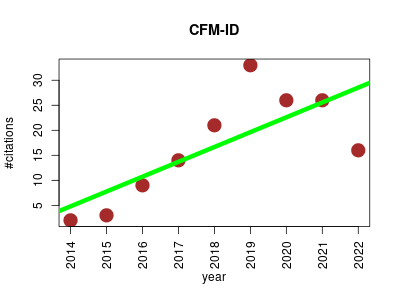 | 26 |
| IMPaLA | Metabolomics Proteomics Systems biology Transcriptomics | IMPaLA is a web tool for the joint pathway analysis of transcriptomics or proteomics and metabolomics data. | 2011-10 | 149 | 12.4 | 1.9 | 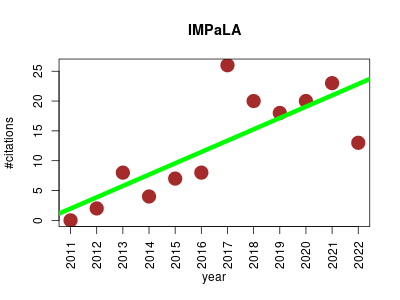 | 31 |
| mzMatch | Data architecture, analysis and design Metabolomics Proteomics experiment | Modular, open source and platform independent data processing pipeline for metabolomics LC/MS data written in the Java language. It was designed to provide small tools for the common processing tasks for LC/MS data. | 2011-04 | 146 | 12.2 | 1.1 | 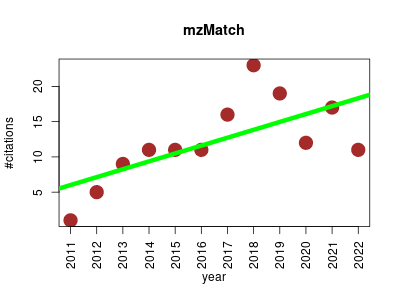 | 12 |
| SMPDB | Endocrinology and metabolism Metabolomics Molecular interactions, pathways and networks Pathology Small molecules | Interactive, visual database containing more than 350 small-molecule pathways found in humans. It is designed specifically to support pathway elucidation and pathway discovery in clinical metabolomics, transcriptomics, proteomics and systems biology. It provides exquisitely detailed, hyperlinked diagrams of human metabolic pathways, metabolic disease pathways, metabolite signaling pathways and drug-action pathways. | 2009-11 | 137 | 9.8 | 1.1 | 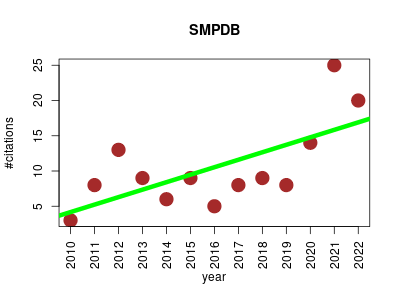 | 17 |
| export2graphlan | Biomarkers Metabolomics Taxonomy | export2graphlan is a conversion software tool for producing both annotation and tree file for GraPhlAn. In particular, the annotation file tries to highlight specific sub-trees deriving automatically from input file what nodes are important. | 2015-01 | 137 | 17.1 | 8.1 | 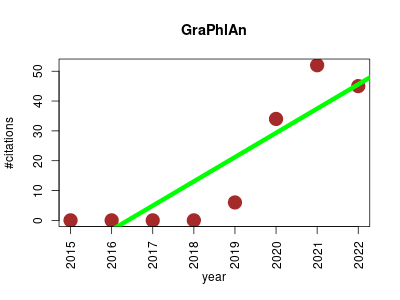 | 6 |
| LipidXplorer | Lipids Metabolomics | LipidXplorer is a software that supports the quantitative characterization of complex lipidomes by interpreting large datasets of shotgun mass spectra. | 2012-01 | 122 | 11.1 | 2.4 | 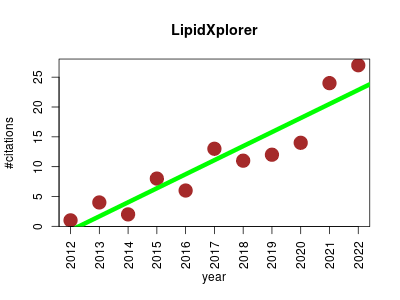 | 13 |
| Galaxy Australia | Bioinformatics Genomics Metabolomics Proteomics | Galaxy Australia is a hosted instance of the Galaxy web-based platform for data intensive biological research, for all Australian researchers. | 2021-01 | 115 | 57.5 | 25.5 | 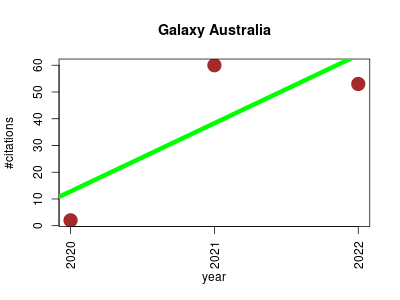 | 2 |
| PRISM 3 | Cheminformatics Metabolomics Structure prediction | PRediction Informatics for Secondary Metabolomes. Prediction of genetically encoded natural product structures based on microbial genomes. It takes a microbial genome sequence as input, identifies biosynthetic gene clusters, and generates combinatorial libraries of structure predictions. | 2017-07 | 113 | 18.8 | 4.8 | 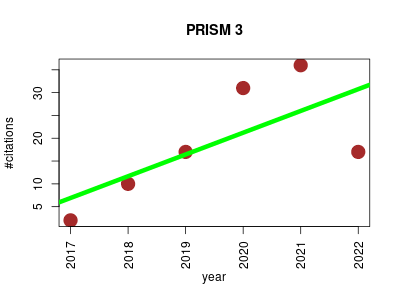 | 38 |
| MZmine | Metabolomics Proteomics Proteomics experiment Small molecules | Toolbox for visualization and analysis of LC-MS data in netCDF or mzXML. | 2005-07 | 107 | 5.9 | -0.2 |  | 20 |
| metaX | Metabolomics Proteomics experiment Sequence analysis | An Automatic and Comprehensive Pipeline for Processing Untargeted Metabolomics Data. | 2017-03 | 101 | 16.8 | 10.5 | 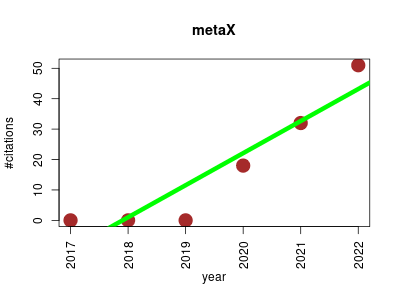 | 5 |
| PolySearch | Genetic variation Metabolomics Molecular interactions, pathways and networks Pathology Small molecules | PolySearch allows users to conduct comprehensive and associative queries, such as given X, find all Y's, where X or Y can be diseases, tissues, cell compartments, gene/protein names, SNPs, mutations, drugs and metabolites. PolySearch also identifies, highlights and ranks abstracts, paragraphs or sentences. | 2008-01 | 100 | 6.7 | -0.1 | 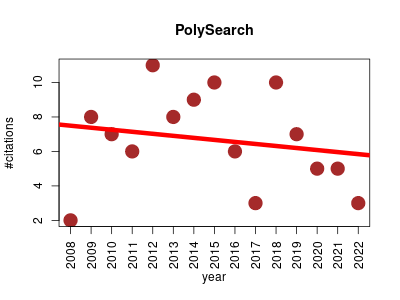 | 22 |
| MS-DIAL | Lipids Metabolomics Proteomics experiment | MS-DIAL was launched as a universal program for untargeted metabolomics that supports multiple instruments (GC/MS, GC/MS/MS, LC/MS, and LC/MS/MS) and MS vendors (Agilent, Bruker, LECO, Sciex, Shimadzu, Thermo, and Waters) | 2020-10 | 94 | 31.3 | 26.0 | 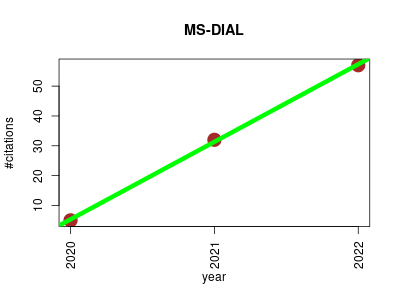 | 14 |
| METASPACE | Imaging Metabolomics Small molecules Zoology | Cloud engine and platform for metabolite annotation for imaging mass spectrometry | 2016-12 | 91 | 13.0 | 4.7 | 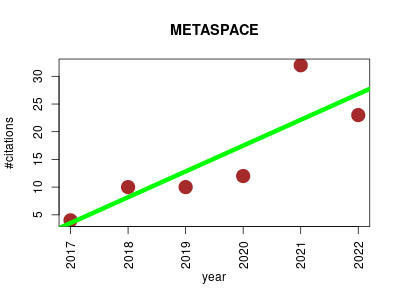 | 13 |
| INMEX | Experimental design and studies Gene expression Metabolomics Microarray experiment Small molecules | Web based tool designed to support meta-analysis of multiple gene-expression data sets, as well as to enable integration of data sets from gene expression and metabolomics experiments. | 2013-01 | 89 | 8.9 | 0.8 | 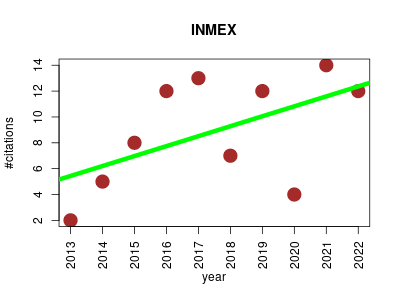 | 37 |
| NOREVA | Metabolomics | NORmalization and EVAluation of MS-based metabolomics data. Enables performance evaluation of various normalization methods from multiple perspectives. In particular, it integrates 5 well-established criteria, each with a distinct underlying theory, to ensure a more comprehensive evaluation than any single criterion. Moreover, it provides various available and popular normalization methods, with a unique feature of allowing quality control based correction sequentially followed by data normalization. | 2017-07 | 89 | 14.8 | 3.5 | 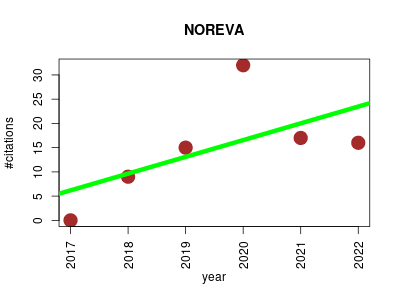 | 11 |
| Sirius | Metabolomics Proteomics Proteomics experiment | Tool for metabolite identification using mass spectrometry. | 2009-01 | 86 | 6.1 | 0.6 | 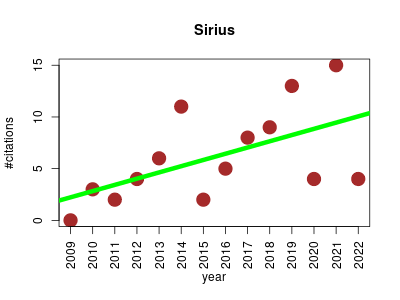 | 12 |
| IMMUNOPHARMACOLOGY | Immunology Metabolomics Parasitology Pharmacology Small molecules | Extending immunopharmacology content and introducing the IUPHAR/MMV Guide to MALARIA PHARMACOLOGY. | 2020-01 | 85 | 28.3 | 2.5 | 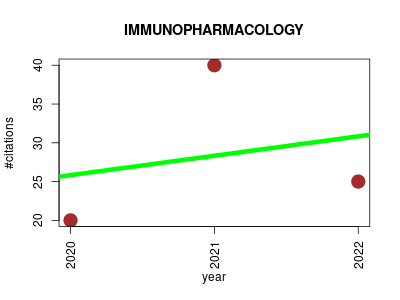 | 25 |
| MassTRIX | Experimental design and studies Mapping Metabolomics Molecular interactions, pathways and networks | With applications in metabolomics and other mass spectrometry studies, MassTRIX is a hypothesis driven approach to the annotation of mass spectrometry data. Data is output in context on a KEGG pathway map. | 2008-01 | 79 | 5.3 | 0.0 | 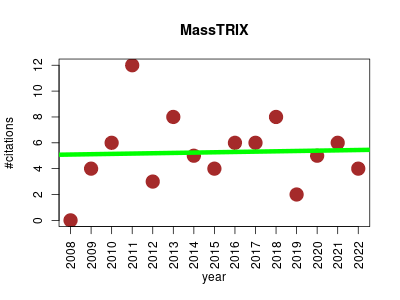 | 9 |
| MetabolomicsTools | Endocrinology and metabolism Metabolomics | Freely-available software tools for metabolomics analysis. | 2017-09 | 76 | 12.7 | 1.9 | 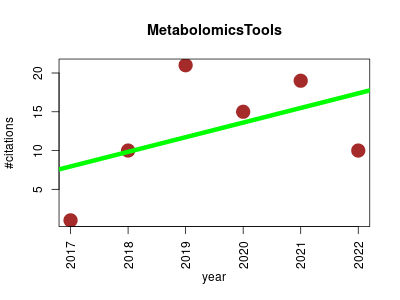 | 26 |
| TargetSearch | Metabolomics | This packages provides a targeted pre-processing method for GC-MS data. | 2009-12 | 74 | 5.3 | 0.4 | 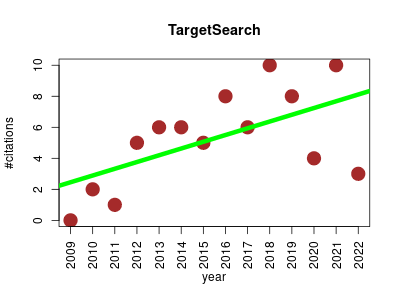 | 4 |
| XCMS Online | Metabolomics Molecular interactions, pathways and networks Systems biology | A systems biology tool for analyzing metabolomic data. It automatically superimposes raw metabolomic data onto metabolic pathways and integrates it with transcriptomic and proteomic data. | 2017-04 | 73 | 12.2 | 1.4 | 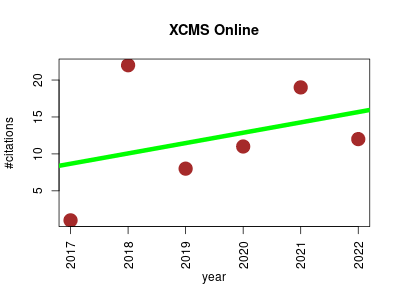 | 17 |
| BridgeDb | Genomics Metabolomics Proteomics Rare diseases | BridgeDb is a framework to map identifiers between various databases. It includes a Java library that provides an API to work with identifier-identifier mapping databases and resources. | 2010-01 | 68 | 5.2 | 0.3 | 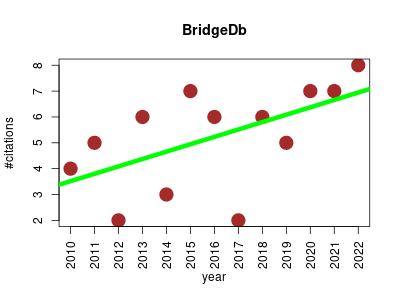 | 6 |
| MaSIF | Machine learning Metabolomics Molecular modelling Protein interactions Protein structural motifs and surfaces | Deciphering interaction fingerprints from protein molecular surfaces using geometric deep learning. | 2020-02 | 67 | 22.3 | 15.5 | 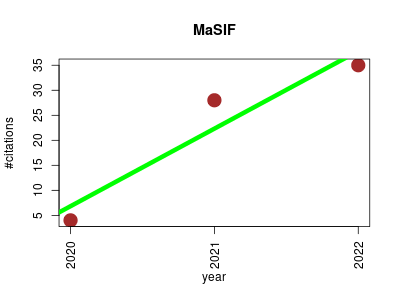 | 18 |
| CANOPUS | Machine learning Metabolomics Plant biology Proteomics experiment Small molecules | CANOPUS is a tool for predicting compound classes directly from MS/MS. Because it does not depend on any database, CANOPUS is suitable for the classification of unknowns. | 2021-04 | 64 | 32.0 | -2.0 | 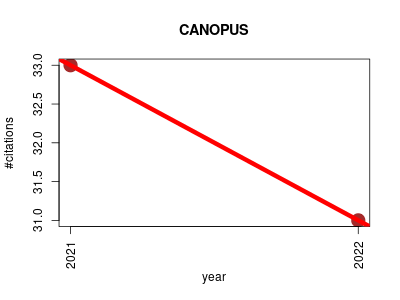 | 16 |
| MBRole | Metabolomics Molecular interactions, pathways and networks | Server that performs functional enrichment analysis of a list of chemical compounds derived from a metabolomics experiment, which allows this list to be interpreted in biological terms. | 2016-07 | 62 | 8.9 | 1.8 | 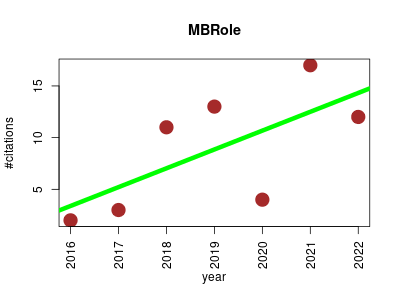 | 8 |
| MetScape 3 | Data visualisation Metabolomics Molecular interactions, pathways and networks | Sparse network modeling and metscape-based visualization methods for the analysis of large-scale metabolomics data. | 2017-05 | 62 | 10.3 | 3.8 | 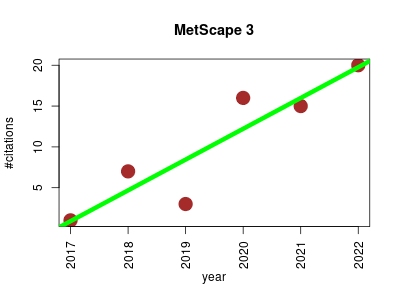 | 12 |
| Babelomics | Data management GWAS study Genomics Metabolomics Proteomics Systems biology Transcriptomics | Integrative platform for the analysis of Transcriptomics, Proteomics and Genomics data with advanced functional profiling. It integrates primary (normalization, calls, etc.) and secondary (signatures, predictors, associations, TDTs, clustering, etc.) analysis tools within an environment that allows relating genomic data and/or interpreting them by means of different functional enrichment or gene set methods. | 2015-01 | 62 | 7.8 | 0.1 | 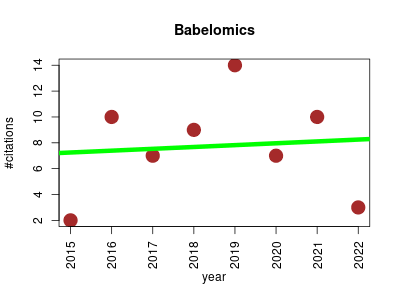 | 4 |
| TPOT | Genotype and phenotype Machine learning Membrane and lipoproteins Metabolomics Workflows | TPOT stands for Tree-based Pipeline Optimization Tool. Consider TPOT your Data Science Assistant. TPOT is a Python Automated Machine Learning tool that optimizes machine learning pipelines using genetic programming. | 2020-01 | 62 | 20.7 | 9.5 |  | 9 |
| xcms | Biological imaging Data visualisation Metabolomics | Framework for processing and visualization of chromatographically separated and single-spectra mass spectral data. The packages enables imports from AIA/ANDI NetCDF, mzXML, mzData and mzML files and preprocesses data for high-throughput, untargeted analyte profiling. | 2010-07 | 61 | 4.7 | 1.0 | 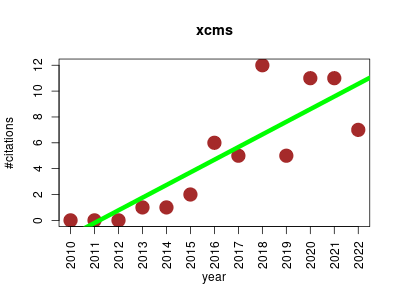 | 1 |
| 3Omics | Data visualisation Data visualisation Human biology Metabolomics Proteomics Systems biology Transcriptomics | A web based systems biology visualization tool for integrating human transcriptomic, proteomic and metabolomic data. | 2013-07 | 56 | 5.6 | 1.0 | 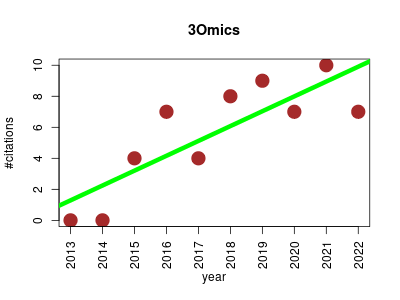 | 31 |
| mmvec | Metabolomics Microbiology Molecular interactions, pathways and networks | Neural networks for estimating microbe-metabolite interactions through their co-occurence probabilities. | 2019-12 | 56 | 14.0 | 7.4 | 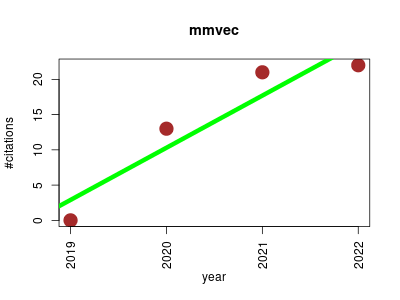 | 13 |
| BiGG Models | Endocrinology and metabolism Metabolomics Molecular interactions, pathways and networks Phylogenetics Small molecules | BiGG Models is a website for browsing gold-standard genome-scale models, multi-strain genome-scale models and expansion across the phylogenetic tree. | 2020-01 | 56 | 18.7 | 1.5 | 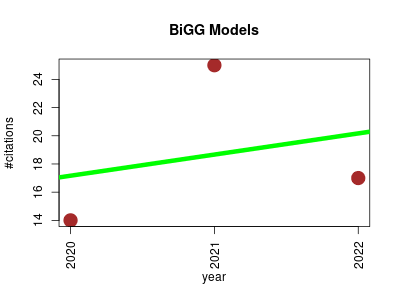 | 17 |
| MetFusion | Metabolomics Proteomics | Approach to combine the knowledge from spectral databases like MassBank with the multitude of candidates generated by fragmenters such as MetFrag. | 2013-03 | 52 | 5.2 | -0.4 | 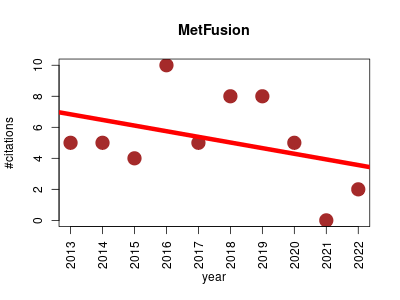 | 13 |
| MetaboMiner | Imaging Metabolomics NMR | Tool which can be used to automatically or semi-automatically identify metabolites in complex biofluids from 2D NMR spectra. It is able to handle both 1H-1H total correlation spectroscopy (TOCSY) and 1H-13C heteronuclear single quantum correlation (HSQC) data. | 2008-11 | 51 | 3.4 | -0.0 | 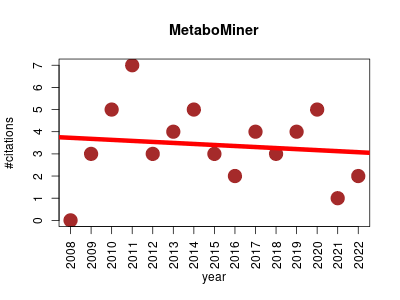 | 14 |
| MetaMapR | Enzymes Metabolomics Molecular interactions, pathways and networks Systems biology | Integrates enzymatic transformations with metabolite structural similarity, mass spectral similarity and empirical associations to generate richly connected metabolic networks. | 2015-01 | 51 | 6.4 | -0.4 | 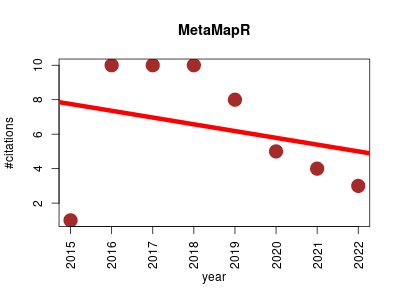 | 18 |
| MathDAMP | Data visualisation Metabolomics Molecular interactions, pathways and networks Small molecules | Facilitates the visualization of differences between metabolite profiles acquired by hyphenated mass spectrometry techniques. Differences are highlighted by applying arithmetic operations to all corresponding signal intensities from whole raw (automatically preprocessed and normalized) datasets on a datapoint-by-datapoint basis. The results are visualized using density plots | 2006-12 | 48 | 2.8 | -0.1 | 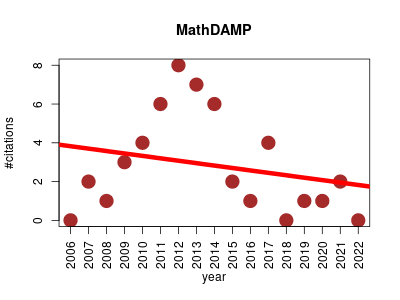 | 8 |
| PeroxisomeDB | Metabolomics Model organisms Model organisms Plants Proteomics | Complete peroxisomal proteome of H. sapiens and S. cerevisiae, as well as 35 newly sequenced eukaryotic genomes. Version 2.0 integrates the peroxisomal metabolome of whole microbody familiy by the new incorporation of the glycosome proteomes of trypanosomatids and the glyoxysome proteome of A. thaliana. The database includes phylogenetic trees and tools to capture kinetic information from Brenda, CheBI and Sabio-RK databases and selected bibliographic references. | 2009-11 | 48 | 3.4 | 0.1 | 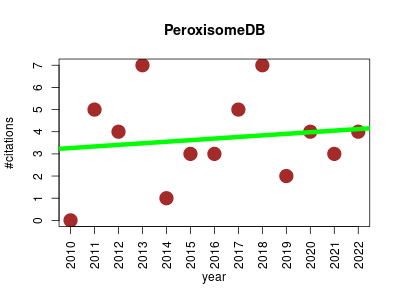 | 6 |
| T3DB | Metabolomics Molecular interactions, pathways and networks Small molecules Toxicology | Unique bioinformatics resource that compiles comprehensive information about common or ubiquitous toxins and their toxin-targets. Each record (ToxCard) contains over 80 data fields providing detailed information on chemical properties and descriptors, toxicity values, protein and gene sequences (for both targets and toxins), molecular and cellular interaction data, toxicological data, mechanistic information and references. | 2009-11 | 47 | 3.4 | 0.2 | 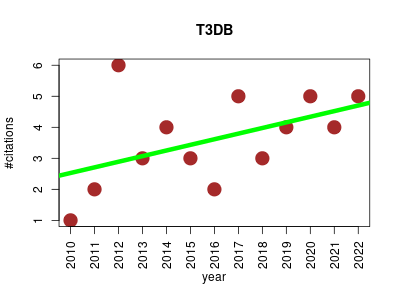 | 12 |
| Lipid Data Analyzer (LDA) | Lipids Metabolomics | The tool enables lipid annotation in high-throughput data derived from chromatography-coupled tandem mass spectrometry. | 2017-12 | 45 | 7.5 | 1.5 | 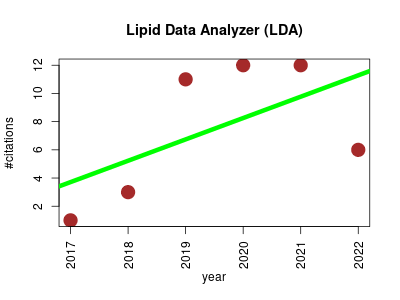 | 6 |
| DeepBGC | Machine learning Metabolomics Small molecules | A deep learning genome-mining strategy for biosynthetic gene cluster prediction | BGC Detection and Classification Using Deep Learning | DeepBGC: Biosynthetic Gene Cluster detection and classification | DeepBGC detects BGCs in bacterial and fungal genomes using deep learning. DeepBGC employs a Bidirectional Long Short-Term Memory Recurrent Neural Network and a word2vec-like vector embedding of Pfam protein domains. Product class and activity of detected BGCs is predicted using a Random Forest classifier | 2019-10 | 43 | 10.8 | 6.1 | 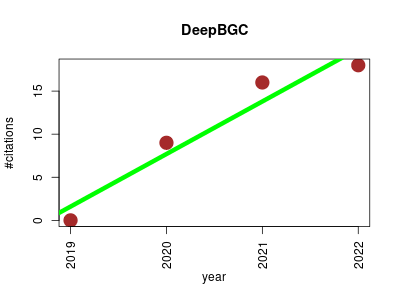 | 15 |
| Metab | Metabolomics Molecular interactions, pathways and networks Systems biology | R package for high-throughput processing of metabolomics data analysed by the Automated Mass Spectral Deconvolution and Identification System (AMDIS) In addition, it performs statistical hypothesis test (t-test) and analysis of variance (ANOVA). Doing so, it considerably speeds up the data mining process in metabolomics and produces better quality results. | 2011-08 | 42 | 3.5 | 0.3 | 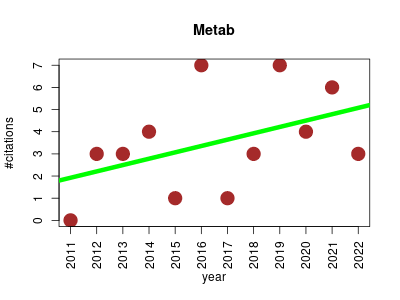 | 7 |
| LipiDex | Lipids Metabolomics Proteomics experiment | A tool for high-confidence LC-MS/MS (Liquid Chromatography-Mass Spectrometry) lipid identification. | 2018-05 | 42 | 8.4 | 2.6 | 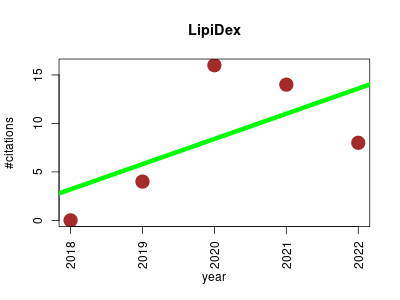 | 7 |
| PRISM | Cheminformatics Endocrinology and metabolism Metabolomics Metagenomics Small molecules | Comprehensive prediction of secondary metabolite structure and biological activity from microbial genome sequences. | 2020-12 | 40 | 13.3 | 8.5 | 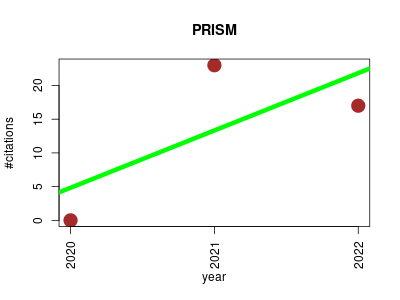 | 15 |
| NMRProcFlow | Metabolomics | An interactive 1D NMR spectra processing tool dedicated to metabolomics. This open source software provides a complete set of tools for processing and visualization of 1D NMR data, the whole within an interactive interface based on a spectra visualization. | 2017-04 | 40 | 6.7 | 2.2 | 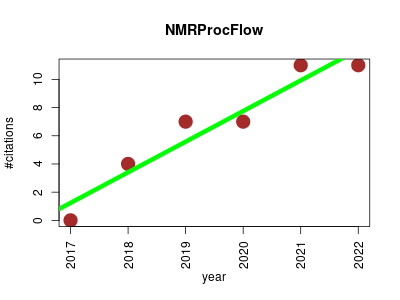 | 7 |
| ANPELA | Metabolomics Metagenomics Proteomics | Analysis and Performance Assessment of Label-free Proteome Quantification. | 2020-03 | 39 | 13.0 | -4.0 | 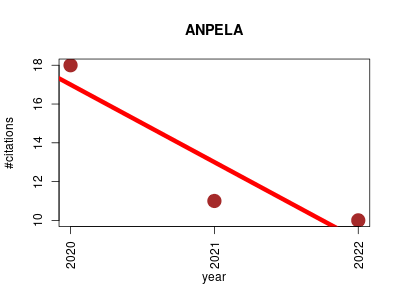 | 5 |
| In silico fragmentation evaluation | Cheminformatics Metabolomics | Comparative analysis of open source in silico fragmentation tools. | 2017-05 | 38 | 6.3 | 1.1 | 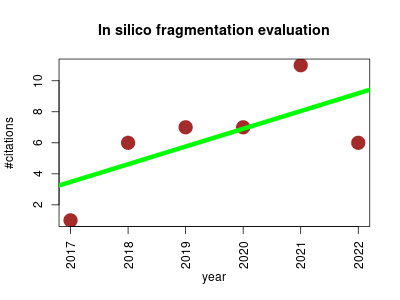 | 7 |
| OSCA | Epigenetics Metabolomics Probes and primers | OSCA (OmicS-data-based Complex trait Analysis) is a software tool for the analysis of complex traits using multi-omics data. | 2019-05 | 38 | 9.5 | 2.8 | 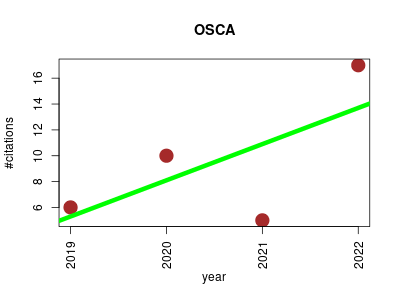 | 8 |
| OpenChrom | Data visualisation Metabolomics Proteomics Proteomics experiment | Open source tool for mass spectrometry and chromatography. | 2010-07 | 37 | 2.8 | 0.7 | 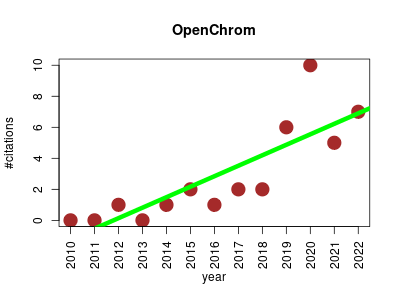 | 3 |
| MeltDB | Metabolomics | Open source framework providing analysis methods for raw GC- and LC-MS metabolome datasets and offers methods to combine the respective results. Flexible tool pipelines allow both the import of pre-processed data as well as the integration of existing open source analysis packages such as XCMS, MassSpecWavelet or metaB. For the identification of metabolites based on mass spectra either the GMD database or user defined libraries in NIST are queried. | 2008-12 | 37 | 2.5 | -0.3 | 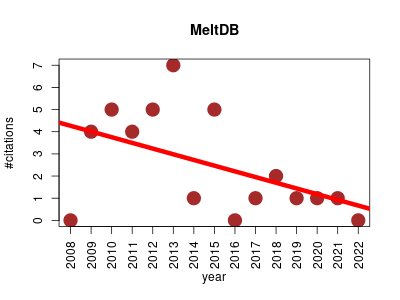 | 9 |
| IMG-ABC | Biotechnology Endocrinology and metabolism Metabolomics Metagenomics Small molecules | An update to the IMG/Atlas of Biosynthetic Gene Clusters Knowledgebase. | 2020-01 | 37 | 12.3 | 2.5 | 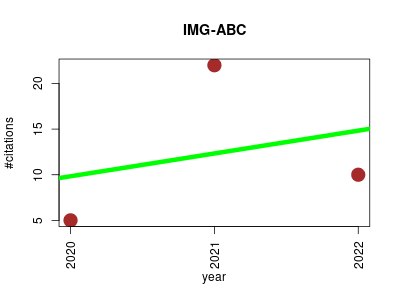 | 18 |
| IDBac | Drug discovery Metabolomics Microbial ecology Microbiology Proteomics | A MALDI bioinformatics platform for analyzing protein and small molecule MALDI-TOF MS data (with a particular focus on mic organism analysis ) | 2018-05 | 35 | 7.0 | 1.6 | 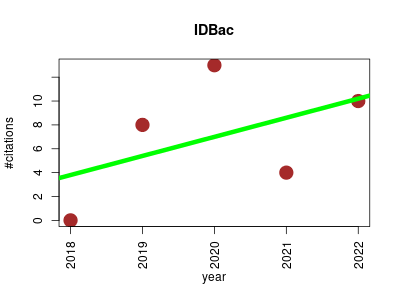 | 9 |
| MPEA | Metabolomics Proteomics experiment Small molecules Studies Systems biology | MPEA is a rapid tool for functional analysis and biological interpretation of metabolic profiling data. In particular, MPEA is designed to be used with data generated by gas chromatography mass spectrometry (GCMS); one of the most prominent analytical methods for metabolic studies and able to quantify hundreds of small molecules from biological extracts in a single run. | 2011-07 | 35 | 2.9 | 0.1 | 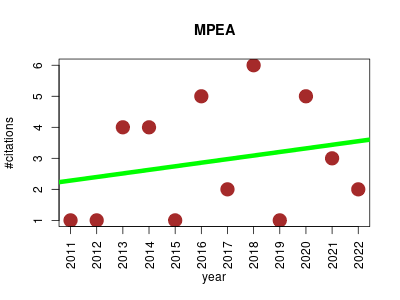 | 13 |
| PAPi | Metabolomics Molecular interactions, pathways and networks | The Pathway Activity Profiling is an R package for predicting the activity of metabolic pathways based solely on a metabolomics data set containing a list of metabolites identified and their respective abundances in different biological samples. It generates hypothesis that improves the final biological interpretation. | 2010-12 | 34 | 2.6 | 0.3 | 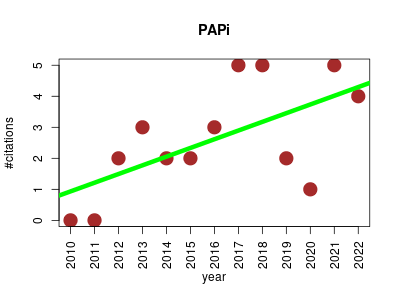 | 6 |
| geoRge | Metabolomics | Analyze untargeted LC/MS data from stable isotope-labeling experiments useing unlabeled and labeled biologically equivalent samples to compare isotopic distributions in the mass spectra. | 2016-01 | 33 | 4.7 | 0.2 | 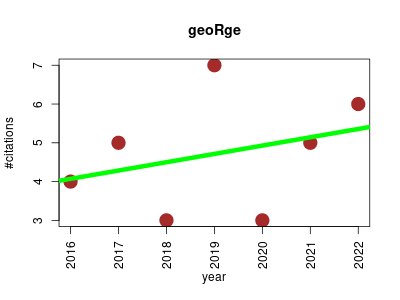 | 5 |
| CEU Mass Mediator | Metabolomics | Tool for looking up metabolites in different databases (Kegg, HMDB, LipidMaps, Metlin and MINE) from experimental masses obtained by mass spectrometry. | 2018-05 | 31 | 6.2 | 1.2 | 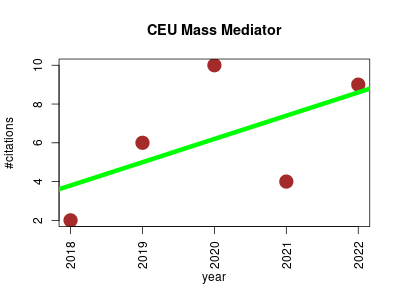 | 1 |
| ProGeM | Epigenetics Gene expression Genotype and phenotype Lipids Metabolomics | A framework for the prioritization of candidate causal genes at molecular quantitative trait loci. | 2019-01 | 31 | 7.8 | 4.1 | 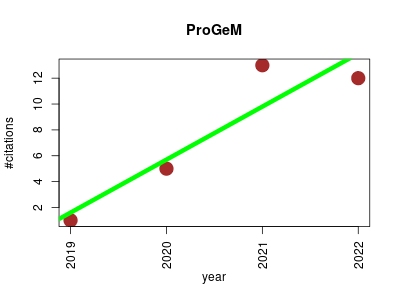 | 5 |
| MAMBO | Data mining Metabolomics Metagenomics Microbial ecology | MAMBO (Metabolomic Analysis of Metagenomes using fBa and Optimization) is an algorithm which assesses and optimizes correlations between genome-scale, constraint-based metabolic models and microbial abundance profiles obtained from shotgun sequence data. | 2018-04 | 30 | 6.0 | 0.6 | 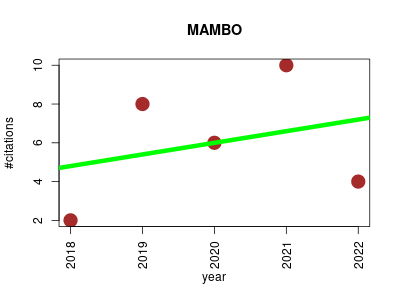 | 10 |
| metaP | Metabolomics | The server automates data analysis for the processing of metabolomics experiments. | 2011-01 | 30 | 2.5 | -0.4 | 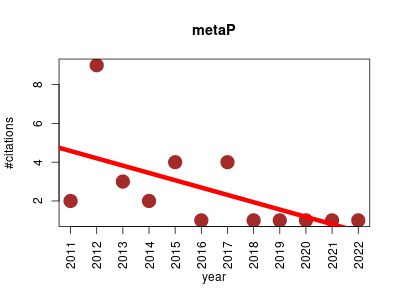 | 10 |
| IsoCor | Fluxomics Metabolomics Systems biology | IsoCor is a scientific software dedicated to the correction of low- and high-resolution mass spectrometry (MS) data for naturally occuring isotopes. IsoCor corrects raw MS data (mass fractions) for naturally-occurring isotopes of all elements and purity of the isotopic tracer. The output of IsoCor is the isotopologue distribution of the molecule (i.e. the relative fractions of molecular entities differing only in the number of isotopic substitutions of the tracer). IsoCor also calculates the mean enrichment (i.e. the mean isotopic content in the molecule) in metabolites. | 2019-11 | 30 | 7.5 | 3.0 | 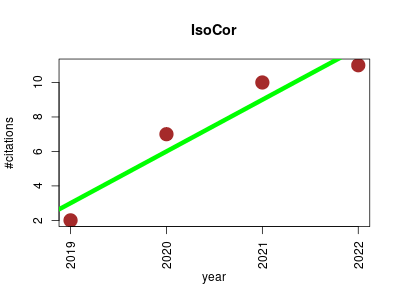 | 2 |
| LFQ-Analyst | Metabolomics Proteomics Proteomics experiment Sequence analysis Small molecules | An Easy-To-Use Interactive Web Platform To Analyze and Visualize Label-Free Proteomics Data Preprocessed with MaxQuant. | 2019-01 | 30 | 7.5 | 2.5 | 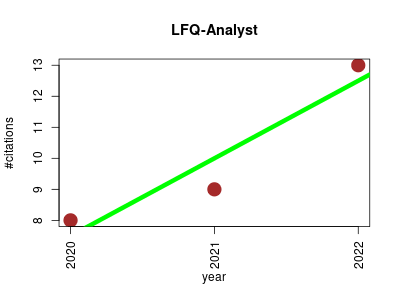 | 3 |
| PathBank | Cell biology Endocrinology and metabolism Metabolomics Molecular interactions, pathways and networks Small molecules | A comprehensive pathway database for model organisms. | 2020-01 | 30 | 10.0 | 4.0 | 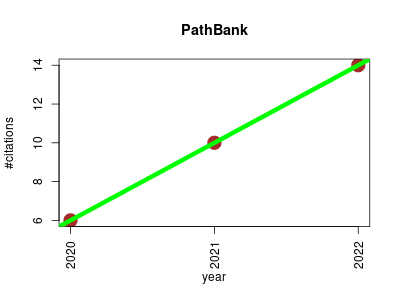 | 10 |
| Deep-DRM | Endocrinology and metabolism Machine learning Metabolomics Pathology Small molecules | Deep-DRM is a computational Method for Identifying Dis-ease-related Metabolites based on Graph Deep Learning Approaches | 2021-07 | 30 | 15.0 | -8.0 | 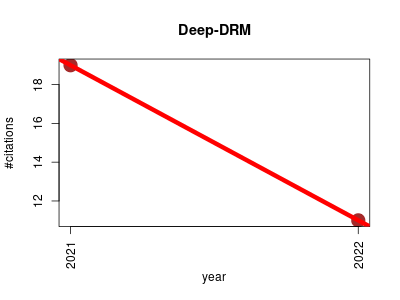 | 3 |
| ProbMetab | Metabolomics Molecular interactions, pathways and networks | ProbMetab is an R package for Bayesian probabilistic annotation of LC–MS-based metabolomics | 2014-05 | 30 | 3.3 | 0.3 | 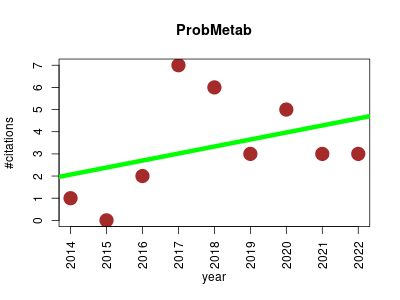 | 10 |
| MSPrep | Metabolomics Systems biology | R package for post processing of metabolomic data. Performs summarization of replicates, filtering, imputation, normalization, generates diagnostic plots and outputs final analytic datasets for downstream analysis. | 2014-01 | 28 | 3.1 | 0.1 | 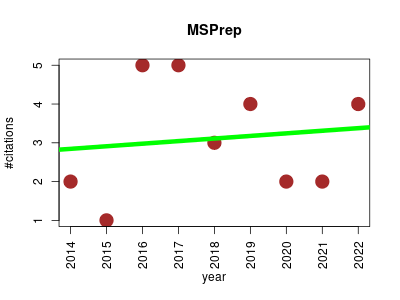 | 6 |
| metaRbolomics | Lipids Metabolomics NMR | The metaRbolomics Toolbox in Bioconductor and beyond | Metabolomics in R and Bioconductor | The term metaRbolomics has been coined for a workshop at the 2016 annual conference of the international Metabolomics society in Dublin, Ireland | Jan Stanstrup, Corey D | 2019-10 | 28 | 7.0 | 2.2 | 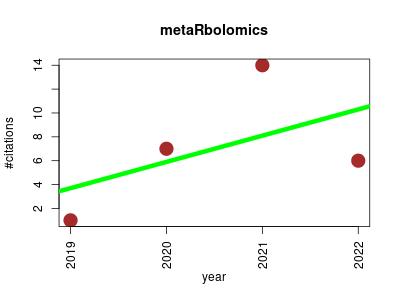 | 11 |
| Blood Exposome Database | Metabolomics Natural language processing Toxicology | Generating the Blood Exposome Database Using a Comprehensive Text Mining and Database Fusion Approach | The exposome represents the sum of all exposures during the life-span of an organism (from chemicals to microbes, viruses, radiation and other sources) | 2019-09 | 28 | 7.0 | 4.2 | 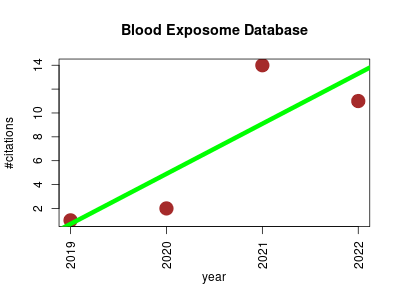 | 8 |
| DeepRiPP | Cell biology Chemistry Machine learning Metabolomics Small molecules | DeepRiPP integrates multiomics data to automate discovery of novel ribosomally synthesized natural products. | 2020-01 | 27 | 9.0 | -1.5 | 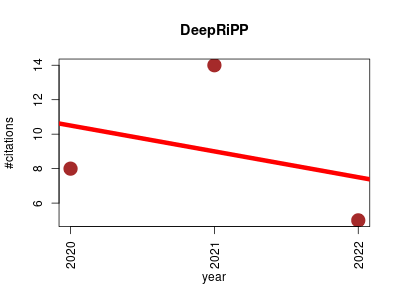 | 15 |
| RMassBank | Metabolomics | Workflow to process tandem MS files and build MassBank records. Functions include automated extraction of tandem MS spectra, formula assignment to tandem MS fragments, recalibration of tandem MS spectra with assigned fragments, spectrum cleanup, automated retrieval of compound information from Internet databases, and export to MassBank records. | 2013-01 | 26 | 2.6 | -0.2 | 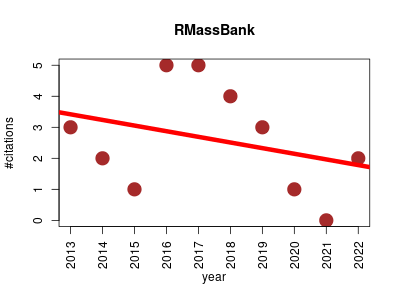 | 6 |
| speaq | Biomarkers Metabolomics NMR | A framework for analyzing automatically raw 1D NMR spectra data. It is also compatible with other NMR frameworks or complementary LC-MS workflows. | 2018-03 | 26 | 5.2 | 2.6 | 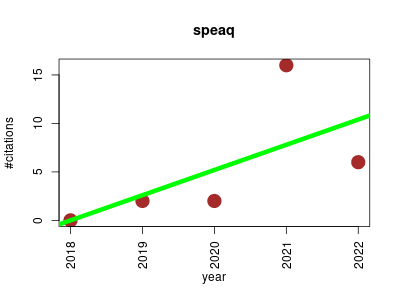 | 8 |
| MetFamily | Metabolomics | Web application designed for the identification of regulated metabolite families. This is possible on the basis of metabolite profiles for a set of MS¹ features as well as one MS/MS spectrum for each MS¹ feature. Group-discriminating MS¹ features are identified using a PCA of metabolite profiles and metabolite families are identified using a HCA of MS/MS spectra. Regulated metabolite families are identified by considering group-discriminating MS¹ features from corporate metabolite families. | 2016-08 | 26 | 3.7 | 0.5 | 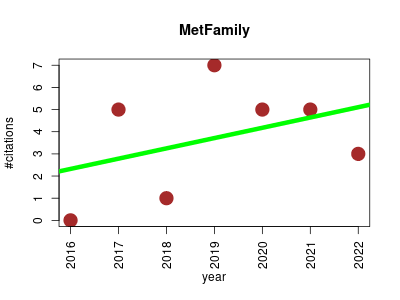 | 5 |
| metabnorm | Metabolomics Systems biology | Mixed model normalization method for metabolomics data. | 2014-08 | 25 | 2.8 | 0.1 |  | 8 |
| LOBSTAHS | Metabolomics Small molecules | Multifunction package for screening, annotation, and putative identification of mass spectral features in large, HPLC-MS lipid datasets. In silico data for a wide range of lipids, oxidized lipids, and oxylipins can be generated from user-supplied structural criteria with a database generation function. It applies these databases to assign putative compound identities to features in any high-mass accuracy dataset that has been processed using xcms and CAMERA. | 2016-07 | 25 | 3.6 | 0.4 | 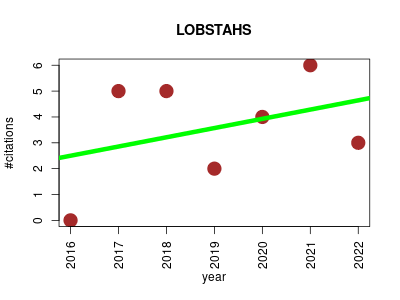 | 7 |
| PhenoMeNal | Metabolomics NMR Phenomics | PhenoMeNal (Phenome and Metabolome aNalysis) is a comprehensive and standardised e-infrastructure that supports the data processing and analysis pipelines for molecular phenotype data generated by metabolomics applications. | 2019-02 | 25 | 6.2 | -0.3 | 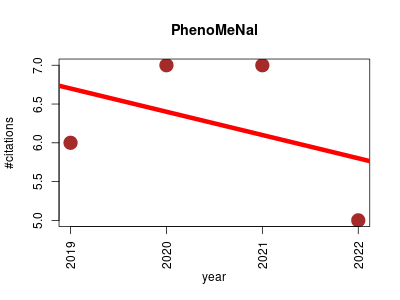 | 6 |
| PiMP | Metabolomics Proteomics experiment Small molecules | PiMP my metabolome: an integrated, web-based tool for LC-MS metabolomics data. | 2017-12 | 25 | 4.2 | 1.1 | 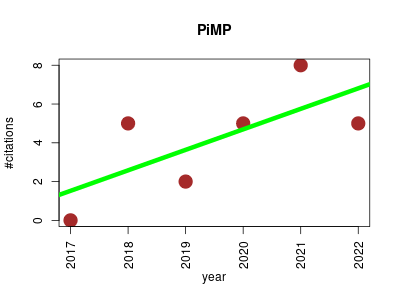 | 1 |
| IEMbase | Biomarkers Medicine Metabolomics Rare diseases | Inborn Errors of Metabolism Knowledgebase (IEMbase) accepts an array of biochemical and clinical symptoms from a user and returns a ranked list of possible IEM disorders that match the input profile. In addition, the system can explain the rationale of its results, suggest possible tests that would assist in narrowing down the differential diagnosis, and provide access to its database of biochemical, molecular, and clinical information if more evidence is desired. | 2018-05 | 24 | 4.8 | 0.5 |  | 10 |
| matTFA | Endocrinology and metabolism Metabolomics Small molecules | Matlab implementation of Thermodynamics-based Flux Analysis. | 2019-01 | 24 | 6.0 | 2.2 | 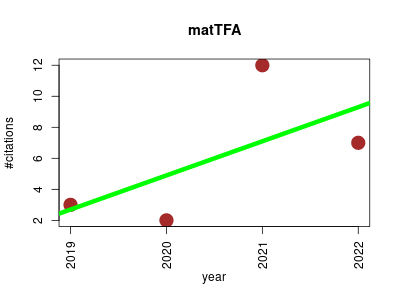 | 5 |
| pyTFA | Endocrinology and metabolism Metabolomics Small molecules | Python package for thermodynamics-based flux analysis. | 2019-01 | 24 | 6.0 | 2.2 |  | 5 |
| CliqueMS | Metabolomics Proteomics experiment Small molecules | Computational tool for annotating in-source metabolite ions from LC-MS untargeted metabolomics data based on a coelution similarity network. | 2019-10 | 24 | 6.0 | 0.6 | 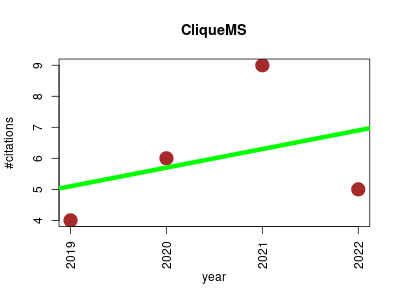 | 5 |
| EcoPrestMet | Drug metabolism Metabolomics Pharmacology | Metabolomics-Driven Exploration of the Chemical Drug Space to Predict Combination Antimicrobial Therapies. | 2019-06 | 23 | 5.8 | 2.5 | 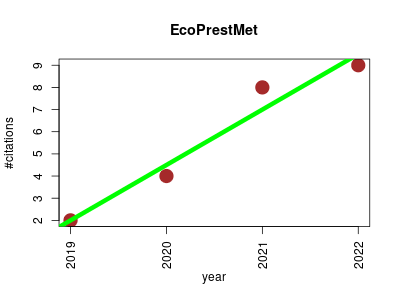 | 11 |
| CNN-T4SE | Machine learning Metabolomics Pharmacology Protein secondary structure Small molecules | Convolutional neural network-based annotation of bacterial type IV secretion system effectors with enhanced accuracy and reduced false discovery. | 2020-09 | 23 | 7.7 | -0.0 | 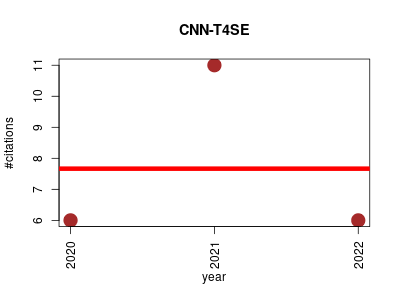 | 4 |
| N-GlycositeAtlas | Metabolomics Protein sites, features and motifs Small molecules | A database resource for mass spectrometry-based human N-linked glycoprotein and glycosylation site mapping | Not a member? Reigist in www.biomarkercenter.org/nglycositeatlas | Database UniProtKB AC/ID Gene Protein Site Peptide Motif Source Year Reference | 2019-09 | 23 | 5.8 | 1.5 | 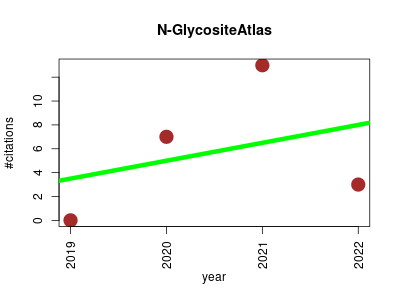 | 6 |
| CCSbase | Biological databases Metabolomics Physics Proteomics experiment Small molecules | CCSbase is an all-in-one web interface for querying the CCS database and accessing the predictive model to support unknown compound identifications. | 2020-03 | 23 | 7.7 | 5.0 | 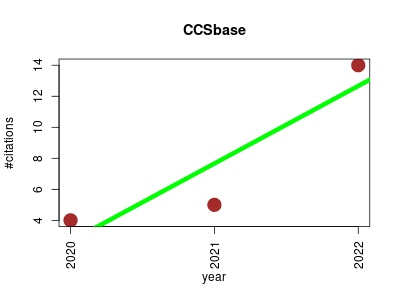 | 2 |
| SpaceM | Endocrinology and metabolism Imaging Metabolomics Small molecules | A method for in situ single-cell metabolomics of cultured cells that integrates microscopy with MALDI-imaging mass spectrometry. | 2021-07 | 23 | 11.5 | 7.0 | 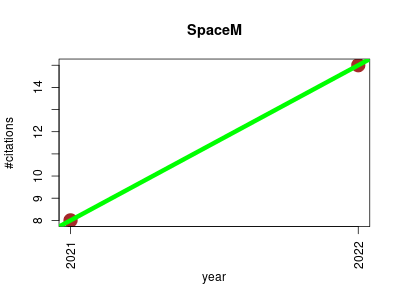 | 13 |
| metaMS | Metabolomics | MS-based metabolomics data processing and compound annotation pipeline. | 2014-09 | 22 | 2.4 | 0.3 | 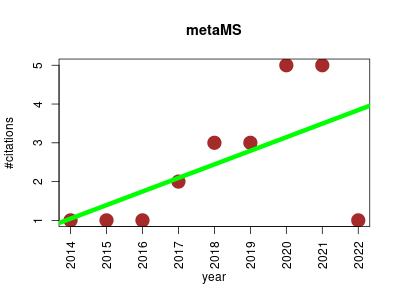 | 8 |
| lipidr | Data mining Lipids Metabolomics Proteomics experiment Workflows | A Software Tool for Data Mining and Analysis of Lipidomics Datasets. | 2020-07 | 22 | 7.3 | 3.5 | 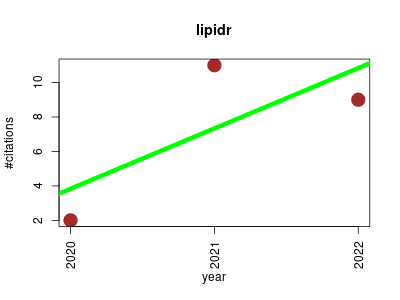 | 5 |
| Yeast MetaboliNER | Literature and language Metabolomics Natural language processing Small molecules | Yeast MetaboliNER is a tool to automatically identify metabolite names in the literature, and associate structures where possible, to define the reported yeast metabolome. | -- | 21 | 0.0 | -0.1 | 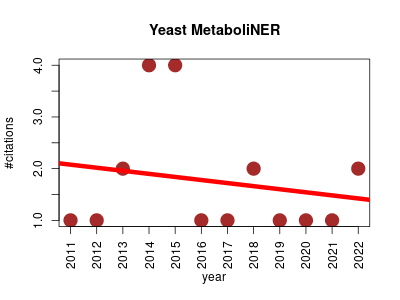 | 6 |
| MMMDB | Metabolomics Molecular interactions, pathways and networks Small molecules | Mouse Multiple Tissue Metabolome Database: metabolomic database containing a collection of metabolites measured from multiple tissues from single mice. The datases are collected using a single instrument and not integrated from literatures, which is useful for capturing the holistic overview of large metabolomic pathway. | 2012-01 | 21 | 1.9 | 0.4 | 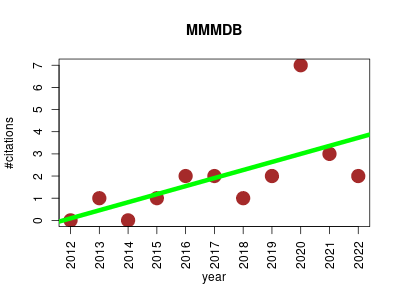 | 6 |
| El-MAVEN | Metabolomics Molecular interactions, pathways and networks Proteomics experiment | HOMEPAGE MISSING! | A Fast, Robust, and User-Friendly Mass Spectrometry Data Processing Engine for Metabolomics | Analysis of large metabolomic datasets is becoming commonplace with the increased realization of the role that metabolites play in biology and pathophysiology. While there are many open-source analysis tools to extract peaks from liquid chromatography-mass spectrometry (LC-MS), gas chromatography-mass spectrometry (GC-MS), and tandem mass spectrometry (LC-MS MS) data, these tools are not very interactive and are suboptimal when a large number of samples are to be analyzed. El-MAVEN is an open-source analysis platform that extends MAVEN and provides fast, powerful, and interactive analysis capabilities especially for datasets containing over 100 samples. The El-MAVEN workflow is easy to use with just four steps from loading data to exporting of the results | 2019-01 | 21 | 5.2 | 2.3 |  | 2 |
| pathway | Endocrinology and metabolism Informatics Metabolomics Molecular interactions, pathways and networks Small molecules | Software for pathway/genome informatics and systems biology. | 2021-01 | 21 | 10.5 | 1.0 | 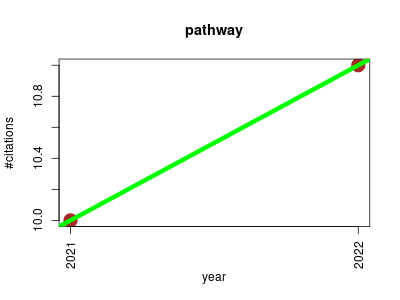 | 3 |
| Exposome Explorer | Biomarkers Metabolomics Nutritional science Oncology Public health and epidemiology | An update incorporating candidate dietary biomarkers and dietary associations with cancer risk. | 2020-01 | 20 | 6.7 | 3.0 | 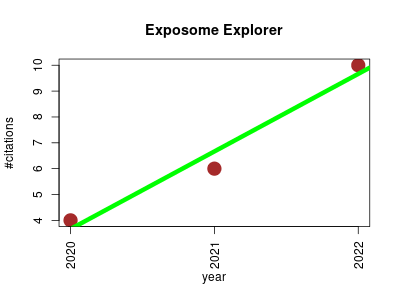 | 10 |
| peakonly | Machine learning Metabolomics Transcription factors and regulatory sites | Deep Learning for the Precise Peak Detection in High-Resolution LC-MS Data. | 2020-01 | 20 | 6.7 | 1.5 | 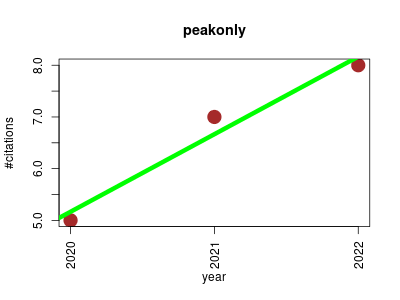 | 11 |
| PMI-DB | Cell biology Metabolomics Protein interactions Proteomics experiment Small molecules | Prediction and collection of protein-metabolite interactions. | -- | 20 | 0.0 | -2.0 | 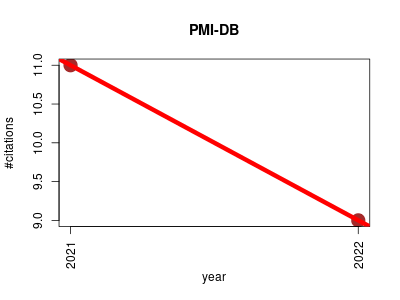 | 2 |
| nmrML converter | Analytical chemistry Data integration and warehousing Metabolomics NMR Ontology and terminology | This tool is used to support the nmrML data standard for nuclear magnetic resonance raw data in metabolomics. It converts native Bruker, Jeol and Agilent vendor formats into the open access nmrML data standard that is used by an ever-so growing number of small molecule NMR repositories. | 2018-01 | 20 | 4.0 | 0.1 | 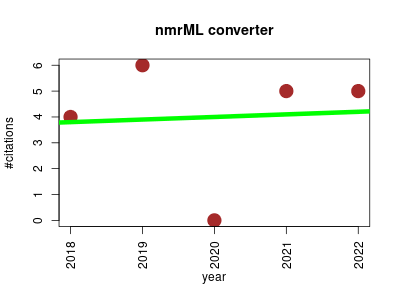 | 10 |
| Warpgroup | Analytical chemistry Metabolomics Protein structure analysis | R package for processing chromatography-mass spectrometry data. It implements: Chromatogram subregion detection, Consensus integration bound determination, Accurate missing value integration. | 2016-01 | 19 | 2.7 | -0.8 | 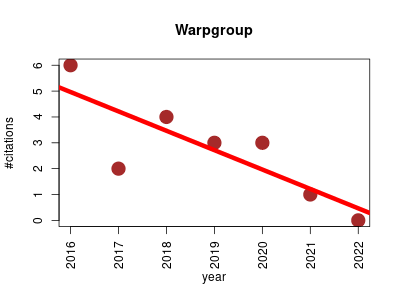 | 6 |
| Retip | Cheminformatics Chemistry Machine learning Metabolomics Proteomics experiment | Retention Time Prediction for Compound Annotation in Untargeted Metabolomics. | 2020-06 | 19 | 6.3 | 4.5 | 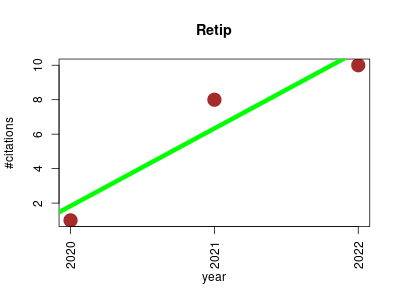 | 4 |
| jMorp | Epigenomics Metabolomics Proteomics experiment Sequencing Small molecules | Large enhancement of multi-omics data resources on the general Japanese population. | 2021-01 | 19 | 9.5 | 9.0 | 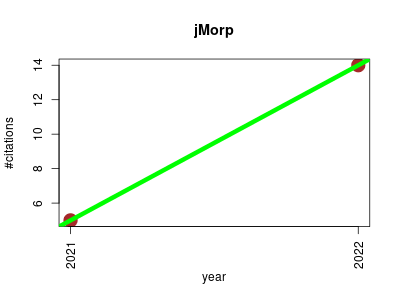 | 3 |
| MetaboDiff | Comparative genomics Metabolomics Sequence analysis | Differential metabolomic analysis. | 2018-10 | 18 | 3.6 | 0.9 | 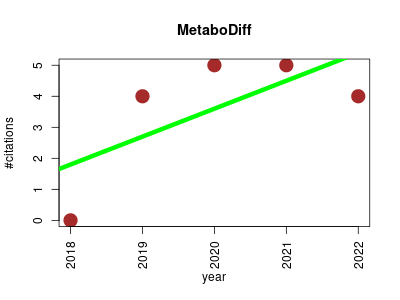 | 5 |
| BIOMEX | Metabolomics Proteomics Sequence analysis Transcriptomics Workflows | an interactive workflow for (single cell) omics data interpretation and visualization. | 2020-01 | 18 | 6.0 | 4.0 | 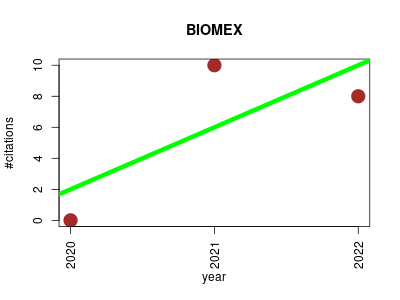 | 3 |
| LungMAP | Anatomy Imaging Metabolomics Ontology and terminology Proteomics | A comparison of alveolar formation and maturation within mouse and human lung. | 2019-10 | 18 | 4.5 | 3.4 | 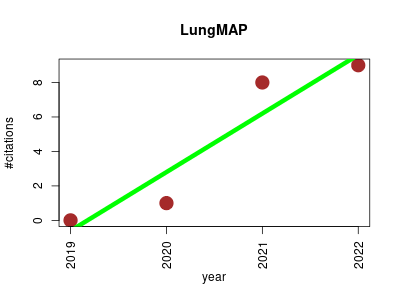 | 7 |
| MS2LDA | Metabolomics Proteomics experiment Small molecules | CORRECT NAME OF TOOL COULD ALSO BE MotifDB | Deciphering complex metabolite mixtures by unsupervised and supervised substructure discovery and semi-automated annotation from MS/MS spectra | Python code for running single file and multi-file LDA | A web application developed in Django+D3 to visualise how topics inferred from Latent Dirichlet Allocation can be used to assist in the unsupervised characterisation of fragmented (LC-MS-MS) metabolomics data | Demo available at www.ms2lda.org (please email us to gain access) | Go to Experiments Quick Decomposition | 25th November 2018: We have introduced many new features to make interpreting Mass2Motifs in MS2LDA.org easier | The last command inserts the gensim lda results into the database | 2019-01 | 17 | 4.2 | 1.5 | 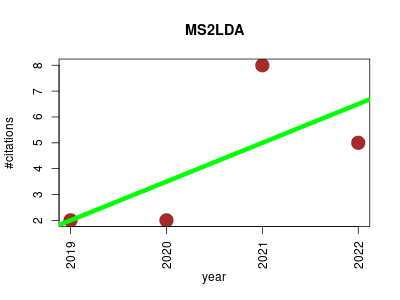 | 7 |
| click-ExM | Cell biology Imaging Lipids Metabolomics Small molecules | Click-ExM is a tool, which integrates click labeling into Expansion microscopy (ExM) to enable a ‘one-stop-shop’ method for nanoscale imaging of various types of biomolecule. By click labeling with biotin and staining with fluorescently labeled streptavidin, a large range of biomolecules can be imaged by the standard ExM procedure normally used for proteins. | 2021-01 | 17 | 8.5 | -3.0 | 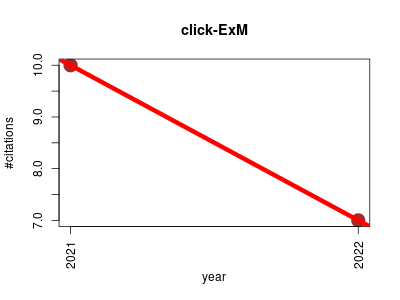 | 7 |
| BRAIN | Analytical chemistry Lipids Metabolomics Proteomics Proteomics experiment Statistics and probability | Package for calculating aggregated isotopic distributions and exact center-masses for chemical substances (in this version composed of C, H, N, O and S). This is an implementation of the BRAIN algorithm. | 2013-02 | 17 | 1.7 | 0.1 | 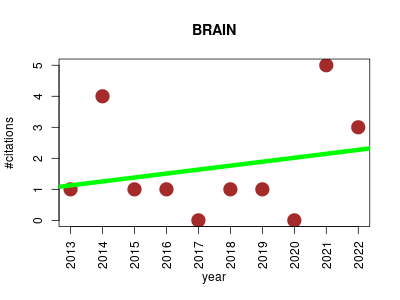 | 0 |
| MetCCS | Metabolomics Proteomics experiment Small molecules | Web server for predicting collision cross-section values of metabolites in ion mobility-mass spectrometry based metabolomics. | 2017-07 | 17 | 2.8 | -0.1 | 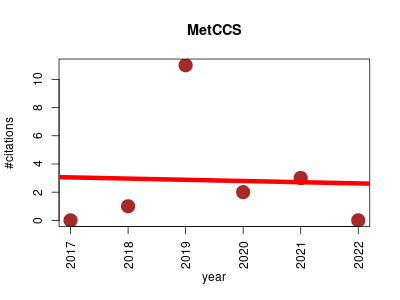 | 5 |
| ChemDistiller | Compound libraries and screening Metabolomics Proteomics experiment Small molecules | Engine for metabolite annotation in mass spectrometry. | 2018-06 | 17 | 3.4 | 1.1 | 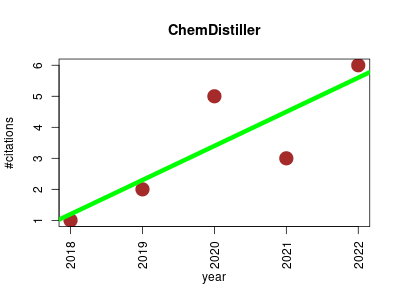 | 3 |
| RegScan | GWAS study Genotype and phenotype Metabolomics | Performance of fast association analysis between allele frequencies and continuous traits. Uses linear regression to estimate marker effects on continuous traits. | 2013-01 | 16 | 1.6 | 0.4 | 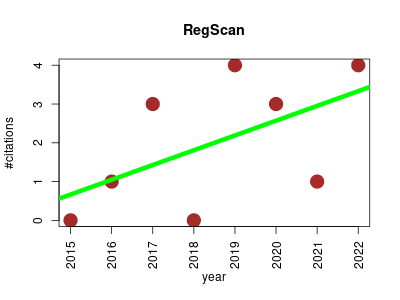 | 3 |
| GeneLab | Epigenomics Metabolomics Metagenomics Proteomics Transcriptomics | Interfaces for the exploration of space omics data. | 2021-01 | 15 | 7.5 | -3.0 | 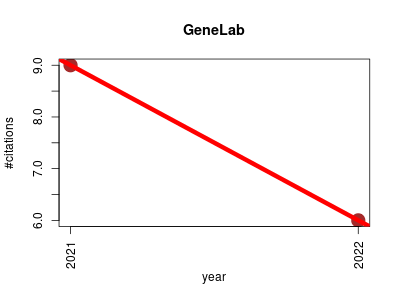 | 3 |
| BMRB | Biophysics Chemistry Metabolomics NMR Small molecules | The Biological Magnetic Resonance Data Bank (BioMagResBank or BMRB) is a repository for data from NMR Spectroscopy on proteins, peptides, nucleic acids, and other biomolecules. | 2020-01 | 15 | 5.0 | 3.0 | 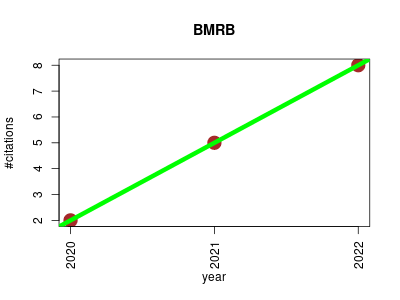 | 7 |
| 'DExSI: Targeted GC-MS Metabolomics' | Metabolomics | An interactive graphical software package which can be used to quantitate isotopologues for a wide variety of metabolites detected by gas chromatography-mass spectrometry. It performs targeted automated metabolite identification, isotopologue mass and abundance determination and natural isotope abundance correction. | 2018-06 | 14 | 2.8 | 0.6 | 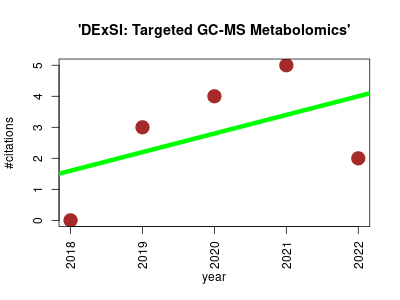 | 0 |
| lmm2met | Metabolomics Pathology Preclinical and clinical studies | Linear mixed-effects modeling to improve clinical metabolomics information. | 2019-01 | 14 | 3.5 | 2.8 | 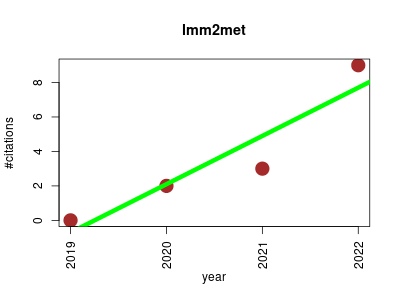 | 3 |
| ICA | Metabolomics Oncology Transcriptomics | Independent Component Analysis for Unraveling the Complexity of Cancer Omics Datasets | ICA-in-Cancer-research-review-materials | Jupyter Notebook containing example of Independent Component Analysis (ICA) accompanying the review in Internal Journal of Molecular Sciences (IJMS) | Independent Component Analysis of BIg Omics Data | BIODICA: Independent Component Analysis for BIg Omics Data | Deconvolution of transcriptome through Immune Component Analysis | BIODICA is a user-friendly pipline for high-performant computation of independent components for omics data, using stability analysis and computing the optimal number of the components from their stabilities, and performing analyses for interpreting the results of ICA application | You can install deconICA from GitHub with: | AN R PACKAGE FOR IDENTIFYING IMMUNE-RELATED SIGNALS IN TRANSCRIPTOME THROUGH DECONVOLUTION OR UNSUPERVISED SOURCE SEPARATION METHODS | 2019-09 | 14 | 3.5 | 0.6 | 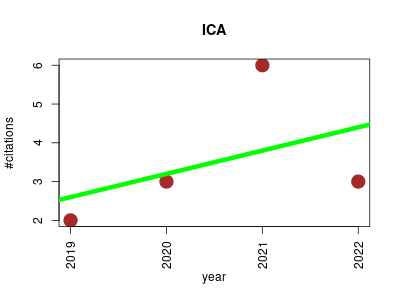 | 4 |
| Binner | Bioinformatics Metabolomics Small molecules | Binner is a Java application for deep annotation of untargeted LC-MS metabolomics data. | 2020-01 | 14 | 4.7 | 0.0 | 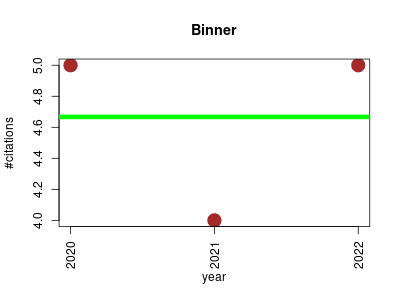 | 5 |
| GLORYx | Drug discovery Endocrinology and metabolism Metabolomics Proteomics experiment Small molecules | Prediction of the Metabolites Resulting from Phase 1 and Phase 2 Biotransformations of Xenobiotics. | 2021-02 | 14 | 7.0 | 2.0 | 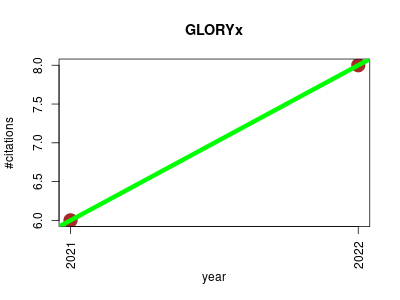 | 3 |
| M2IA | Endocrinology and metabolism Metabolomics Microbial ecology Molecular interactions, pathways and networks Small molecules | A web server for microbiome and metabolome integrative analysis. | 2020-06 | 14 | 4.7 | 1.5 | 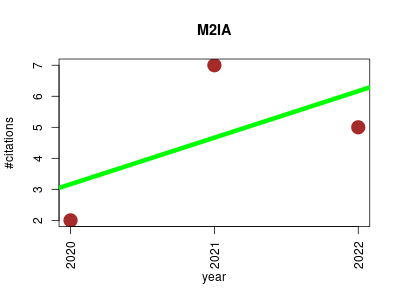 | 3 |
| MetaClean | Machine learning Metabolomics Proteomics Proteomics experiment Small molecules | A machine learning-based classifier for reduced false positive peak detection in untargeted LC-MS metabolomics data. | 2020-11 | 14 | 4.7 | 2.5 | 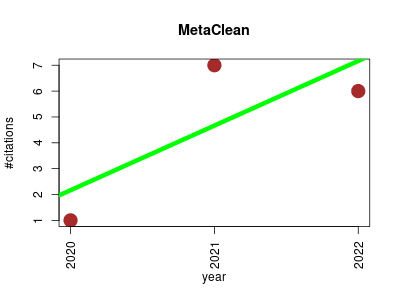 | 4 |
| qnr | DNA Machine learning Metabolomics Sequence analysis | Python implementation of a method for searching large metagenomic dataset to identify qnr fluoroquinolone antibiotic resistance genes. | 2012-12 | 13 | 1.2 | -0.0 | 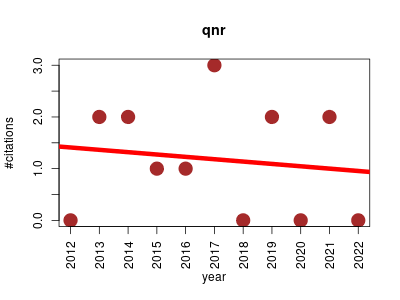 | 4 |
| RaMP | Data management Gene expression Metabolic pathways Metabolomics | A Comprehensive Relational Database of Metabolomics Pathways for Pathway Enrichment Analysis of Genes and Metabolites. | 2018-03 | 13 | 2.6 | -0.4 | 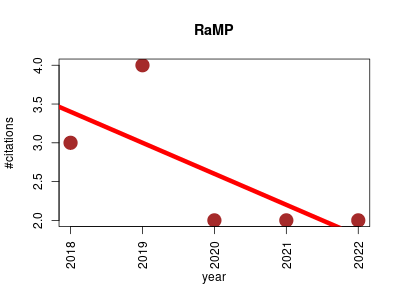 | 7 |
| MFSearcher | Metabolomics | RESTful web service for high-throughput prediction of elemental compositions from accurate mass values detected by high-resolution mass spectrometers. | 2013-01 | 13 | 1.3 | 0.2 | 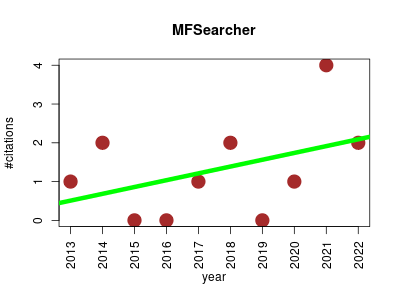 | 2 |
| NERDD | Drug discovery Drug metabolism Endocrinology and metabolism Metabolomics Small molecules | A web portal providing access to in silico tools for drug discovery. | 2020-02 | 13 | 4.3 | 3.5 | 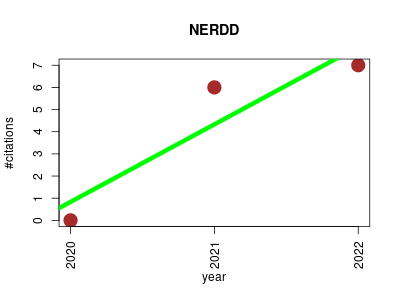 | 1 |
| Maltcms | Algorithms Bioinformatics Metabolomics Pipelines Proteomics | Modular Application Toolkit for Chromatography Mass-Spectrometry is an application framework mainly suited for developers working in the domain of bioinformatics for metabolomics and proteomics. Its aim is to provide reusable, efficient datastructures, abstracting from the various low-level data-formats like netcdf, mzXML, mzData and mzML and providing consistent access to data features like mass spectra, chromatograms and metadata. | 2009-08 | 12 | 0.9 | -0.1 | 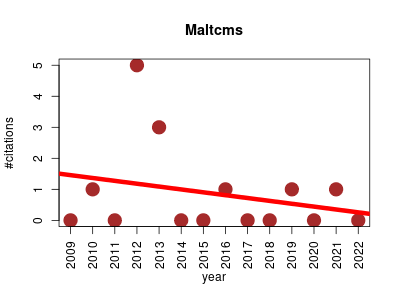 | 2 |
| AMON | Data integration and warehousing Data quality management Metabolomics Microbial ecology Small molecules | annotation of metabolite origins via networks to integrate microbiome and metabolome data. | 2019-11 | 12 | 3.0 | 1.8 | 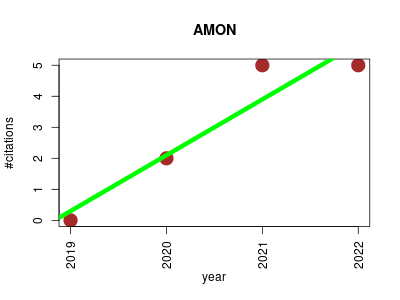 | 6 |
| BWMR | GWAS study Genotype and phenotype Metabolomics Small molecules Statistics and probability | Bayesian weighted Mendelian randomization for causal inference based on summary statistics. | 2020-03 | 12 | 4.0 | 2.0 | 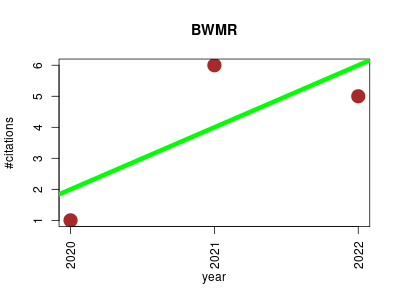 | 2 |
| CycloNovo | Metabolomics Microbial ecology Proteomics experiment Small molecules | De Novo Peptide Sequencing Reveals Many Cyclopeptides in the Human Gut and Other Environments. | 2020-01 | 12 | 4.0 | 0.5 | 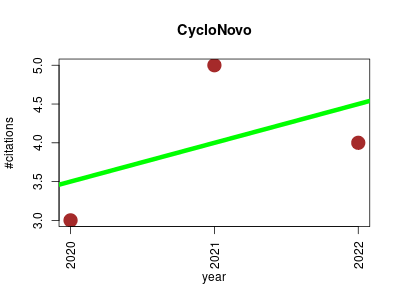 | 3 |
| DecoMetDIA | Analytical chemistry Metabolomics Proteomics experiment | Deconvolution of Multiplexed MS/MS Spectra for Metabolite Identification in SWATH-MS-Based Untargeted Metabolomics | DecoMetDIA was developed to process SWATH-MS based data for metabolomics. | 2019-09 | 12 | 3.0 | 0.4 | 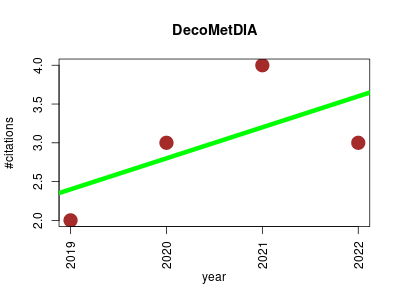 | 5 |
| LDB | Chemistry Metabolomics Molecular biology Proteomics experiment Small molecules | A database of high-resolution MS/MS spectra for lichen metabolites. | 2019-12 | 12 | 3.0 | 1.4 | 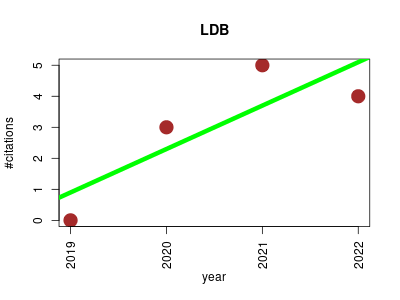 | 3 |
| Real-time | Endocrinology and metabolism Metabolomics Nutritional science Physiology Proteomics experiment | Real-time is a health monitoring through urine metabolomics. | -- | 12 | 0.0 | 2.0 | 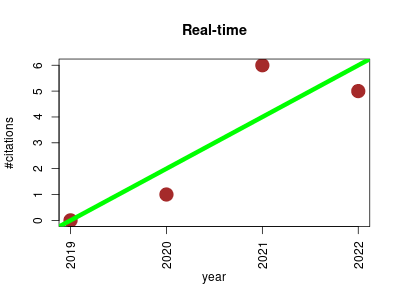 | 5 |
| SF-Matching | Machine learning Metabolomics Molecular biology Proteomics experiment Small molecules | SF-Matching(SubFragment-Matching) is a machine-learning based approach to predict compounds from tandem mass spectra. | 2020-02 | 12 | 4.0 | -0.5 | 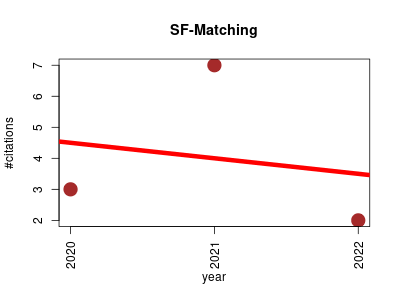 | 5 |
| MSK-KP | Biobank Endocrinology and metabolism Epigenetics Genomics Metabolomics | Making Omics Data Useful to the Broader Scientific Community. | 2020-09 | 12 | 4.0 | 1.5 | 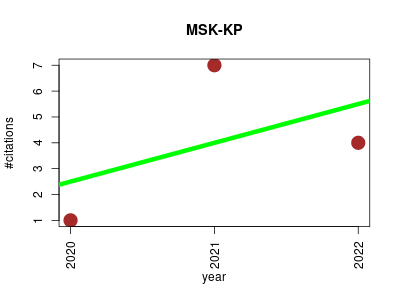 | 7 |
| MEBS | Genomics Metabolomics Metagenomics Molecular interactions, pathways and networks | Software platform to evaluate large (meta)genomic collections according to their metabolic machinery: unraveling the sulfur cycle. | 2017-11 | 12 | 2.0 | 0.5 | 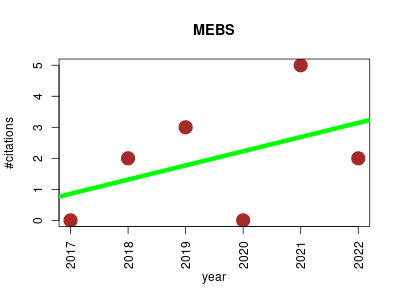 | 0 |
| AssayR | Metabolomics Molecular interactions, pathways and networks Proteomics experiment | Simple Mass Spectrometry Software Tool for Targeted Metabolic and Stable Isotope Tracer Analyses. | 2017-09 | 12 | 2.0 | 0.5 | 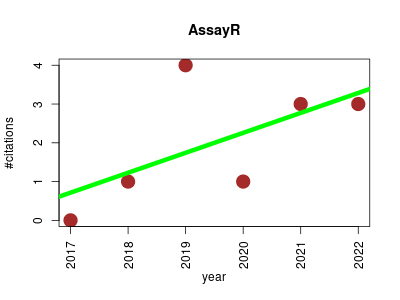 | 1 |
| Lilikoi | Metabolomics Molecular interactions, pathways and networks Small molecules | R package for personalized pathway-based classification modeling using metabolomics data. | 2018-12 | 11 | 2.2 | 0.5 | 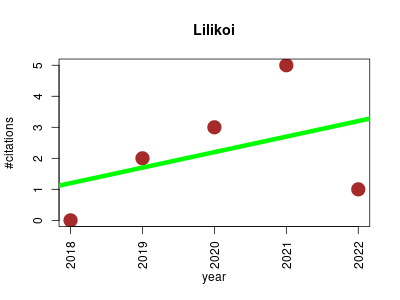 | 5 |
| NaPLeS | Compound libraries and screening Metabolomics Small molecules | A natural products likeness scorer-web application and database | NaPLeS - Natural Product Likeness Score calculator | NaPLeS is an open source web application based fully on open data. Source and installation instructions are available at GitHub. Please submit bug reports, feature requests and general issues through the issues tracker at GitHub. NaPLeS is developed and maintained by the Steinbeck group at the University Friedrich-Schiller in Jena, Germany | 2019-01 | 11 | 2.8 | 1.1 | 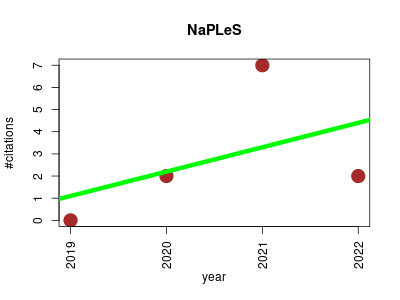 | 4 |
| AutoTuner | Endocrinology and metabolism Metabolomics Proteomics experiment | Autotuner is a used to identify proper parameters during metabolomics data processing. | 2020-04 | 11 | 3.7 | 1.0 | 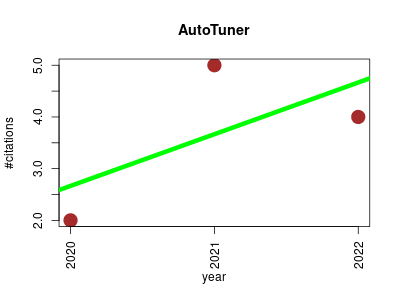 | 5 |
| ICBM-OCEAN | Chemistry Medicine Metabolomics Pharmacology Proteomics experiment | Processing Ultrahigh-Resolution Mass Spectrometry Data of Complex Molecular Mixtures. | 2020-05 | 11 | 3.7 | 3.5 | 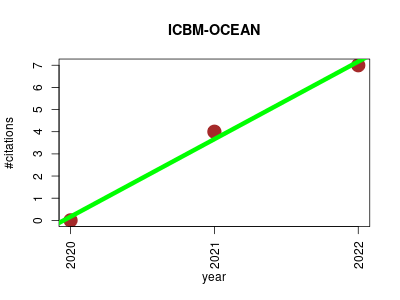 | 1 |
| IP4M | Data mining Metabolomics Microbial ecology Proteomics experiment Workflows | An integrated platform for mass spectrometry-based metabolomics data mining. | 2020-10 | 11 | 3.7 | 3.5 | 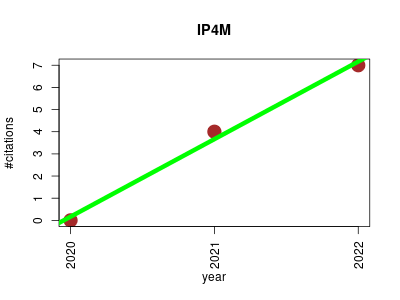 | 3 |
| LPPtiger | Lipids Metabolomics Proteomics experiment Structure analysis | lipidome-specific prediction and identification of oxidized phospholipids from LC-MS datasets. | 2017-12 | 11 | 1.8 | 0.7 | 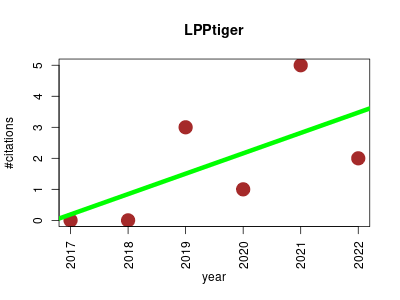 | 6 |
| VSClust | Data mining Genomics Metabolomics Omics Proteomics Statistics and probability | Feature-based variance-sensitive clustering of omics data. Optimizes cluster assignment by taking into account individual feature variance . | 2018-09 | 11 | 2.2 | 0.5 | 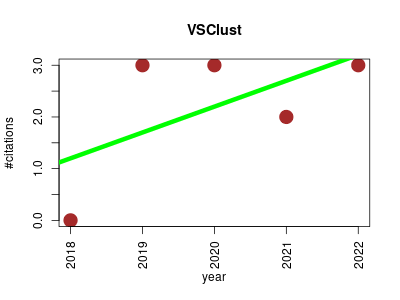 | 0 |
| iPAVS | Endocrinology and metabolism Metabolomics Molecular interactions, pathways and networks Pathology | integrated Pathway Resources, Analysis and Visualization System: provides a collection of highly-structured manually curated human pathway data, it also integrates biological pathway information from several public databases and provides several tools to manipulate,filter, browse, search, analyze, visualize and compare the integrated pathway resources. | 2012-01 | 10 | 0.9 | -0.2 | 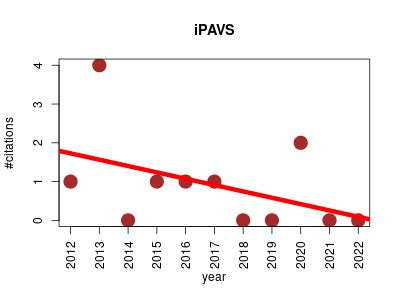 | 2 |
| MetaboSignal | Genetics Metabolomics Molecular interactions, pathways and networks | R package that allows merging, analyzing and customizing metabolic and signaling KEGG pathways. It is a network-based approach designed to explore the topological relationship between genes (signaling or enzymatic genes) and metabolites, representing a powerful tool to investigate the genetic landscape and regulatory networks of metabolic phenotypes. | 2017-01 | 10 | 1.7 | -0.0 | 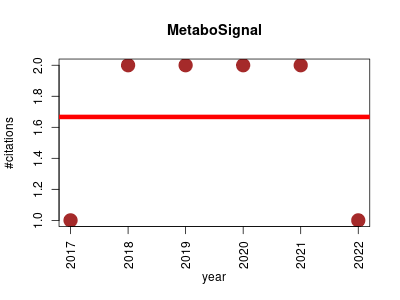 | 3 |
| ROIMCR | Biomarkers Lipids Metabolomics | Powerful analysis strategy for LC-MS metabolomic datasets. | 2019-05 | 10 | 2.5 | 0.8 | 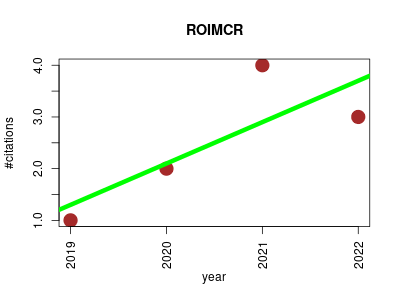 | 6 |
| COLMARm | MRI Metabolomics NMR | Automated Tools for the Analysis of 1D-NMR and 2D-NMR Spectra | COLMARm 13C-1H HSQC, HSQC-TOCSY and TOCSY Query and Verification | Visit a step by step how-to with figures here | Bingol, K.; Li, DW.; Zhang, B.; Brüschweiler, R.; Comprehensive metabolite identification strategy using multiple two-dimensional NMR spectra of a complex mixture implemented in the COLMARm web server; Anal. Chem; 2016, 88, 12411-12418; | 2019-01 | 10 | 2.5 | 1.2 | 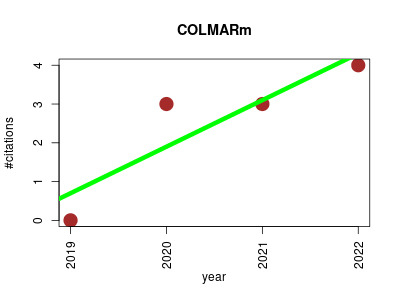 | 6 |
| jmzTab | Metabolomics Proteomics | Provides reading and writing capabilities, as well as supporting the validation of mzTab and the conversion of PRIDE XML and mzIdentML files to mzTab. | 2014-01 | 9 | 1.0 | -0.2 | 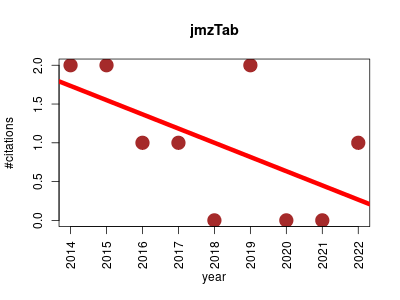 | 2 |
| AspWood | Carbohydrates Metabolomics Plant biology | Whole-genome annotation and functional analyses based on RNA expression data. | 2019-01 | 9 | 2.2 | 0.5 | 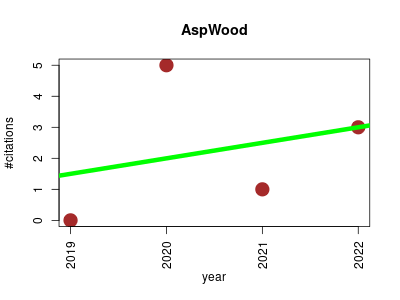 | 2 |
| FOBI | Biomarkers Metabolomics Nutritional science Ontology and terminology Small molecules | FOBI (Food-Biomarker Ontology) an ontology to represent food intake data and associate it with metabolomic data. It is composed of two interconnected sub-ontologies. One is a “Food Ontology” consisting of raw foods and prepared foods while the second is a “Biomarker Ontology” containing food intake biomarkers classified by their chemical classes. | 2020-01 | 9 | 3.0 | 1.5 | 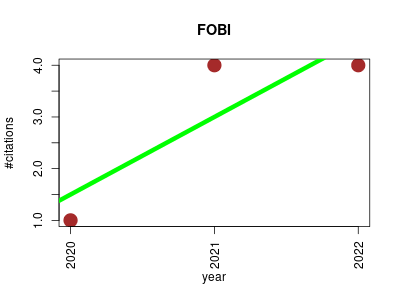 | 4 |
| gutMGene | Metabolomics Metagenomic sequencing Metagenomics Microbial ecology Small molecules | A comprehensive database for target genes of gut microbes and microbial metabolites. | 2022-01 | 9 | 9.0 | 0.0 |  | 1 |
| M-path | Metabolomics Molecular interactions, pathways and networks Systems biology | Calculates random combinations of a wide variety of reactions to list potential metabolic routes from initial to target compounds efficiently, which could be a heuristic approach in designing potential metabolic pathways in practical applications. | 2015-03 | 8 | 1.0 | 0.0 | 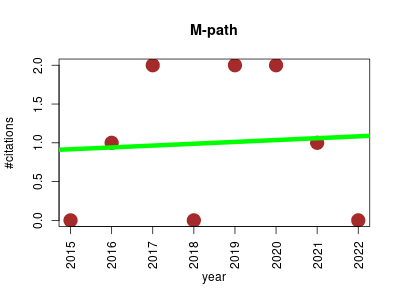 | 4 |
| MetiTree | Metabolomics | Repository, in a prototype stage, of mass spectra of small chemical compounds for life science (2000 Da). The database will contain MSn data acquired on different platforms by different research groups. MSn data will be annotated with chemical information about the fragments. | 2012-10 | 8 | 0.7 | -0.0 | 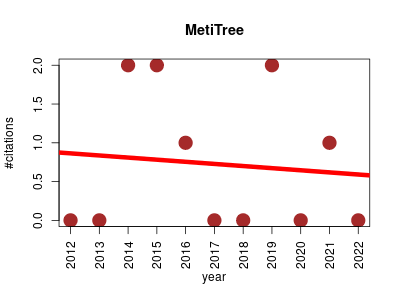 | 3 |
| FELLA | Metabolomics | Enrichment of metabolomics data using KEGG entries. Given a set of affected compounds, FELLA suggests affected reactions, enzymes, modules and pathways using label propagation in a knowledge model network. The resulting subnetwork can be visualised and exported. | 2017-12 | 8 | 1.3 | 0.2 | 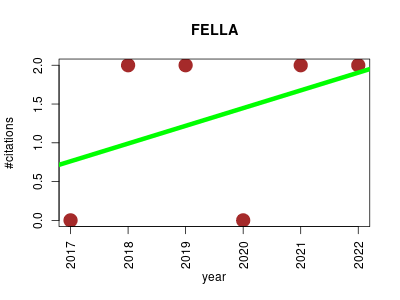 | 2 |
| SpiderMass | Metabolomics | Semantic chemical database generation, metabolite identification and de novo formula generation. | 2015-01 | 8 | 1.0 | -0.0 | 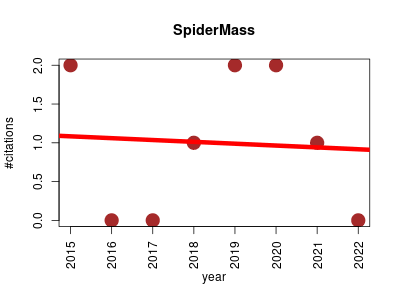 | 0 |
| COMSPARI | Metabolomics Proteomics Proteomics experiment | Compares two datasets in netCDF or ASCII format. | 2004-01 | 8 | 0.4 | -0.1 | 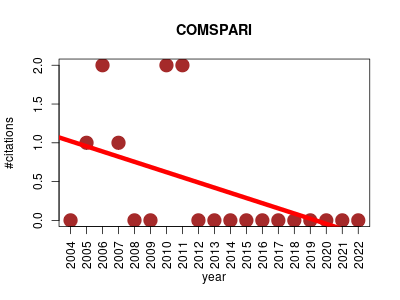 | 3 |
| autoGCMSDataAnal | Metabolomics Plant biology Software engineering | A comprehensive automatic data analysis strategy for gas chromatography-mass spectrometry based untargeted metabolomics. | 2020-04 | 8 | 2.7 | 0.5 | 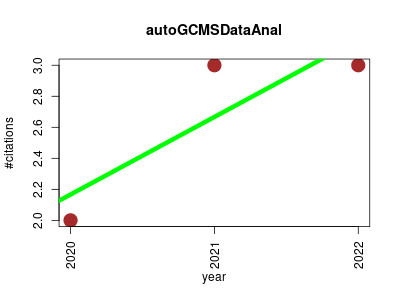 | 3 |
| InterSpin | Metabolomics NMR Small molecules | Integrated Supportive Webtools for Low- and High-Field NMR Analyses Toward Molecular Complexity | Description SpinLIMS Contact Others | Integrated supportive webtools for low- and high-field NMR analysis toward molecular complexities | SENsitivity improvement with Spectral Integration | 2019-02 | 8 | 2.0 | 0.4 | 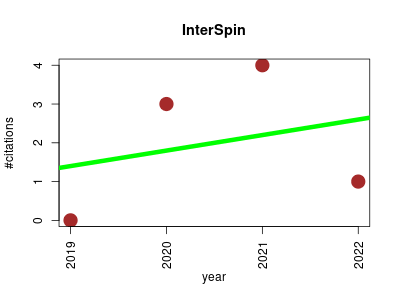 | 3 |
| Medusapy | Endocrinology and metabolism Machine learning Metabolomics Molecular interactions, pathways and networks Small molecules | Software to build and analyze ensembles of genome-scale metabolic network reconstructions. | 2020-04 | 8 | 2.7 | 2.0 | 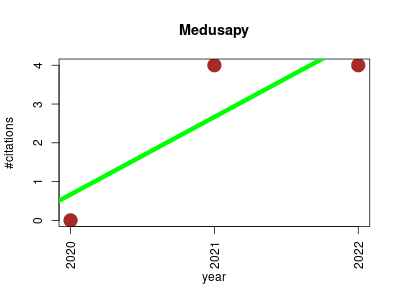 | 2 |
| StanDep | Endocrinology and metabolism Enzymes Mapping Metabolomics Transcriptomics | Capturing transcriptomic variability improves context-specific metabolic models. | 2020-05 | 8 | 2.7 | 1.0 | 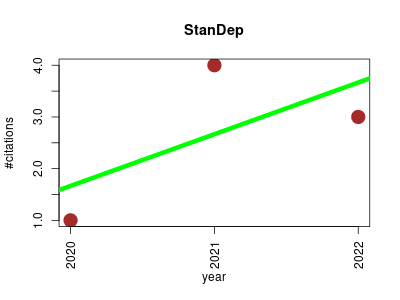 | 3 |
| MOG | Gene expression Metabolomics Microarray experiment Oncology RNA-Seq | A workbench for interactive exploratory data analysis of large expression datasets. | 2020-02 | 8 | 2.7 | -0.0 | 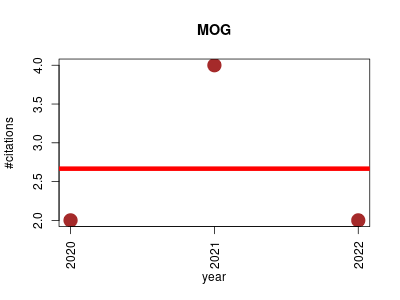 | 2 |
| MMEASE | Metabolomics NMR Proteomics Proteomics experiment Small molecules | Online meta-analysis of metabolomic data by enhanced metabolite annotation, marker selection and enrichment analysis. | 2021-02 | 8 | 4.0 | 0.0 | 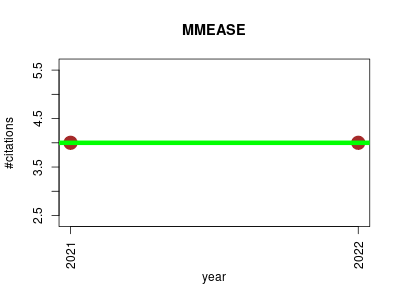 | 3 |
| TeaCoN | Agricultural science Gene expression Metabolomics Plant biology RNA-Seq | a database of gene co-expression network for tea plant (Camellia sinensis). | 2020-07 | 8 | 2.7 | 0.5 | 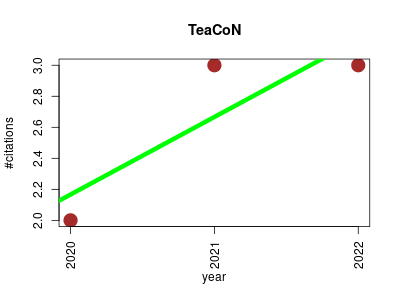 | 1 |
| OffsampleAI | Data mining Imaging Machine learning Metabolomics Proteomics experiment | Artificial intelligence approach to recognize off-sample mass spectrometry images. | 2020-04 | 8 | 2.7 | -1.0 | 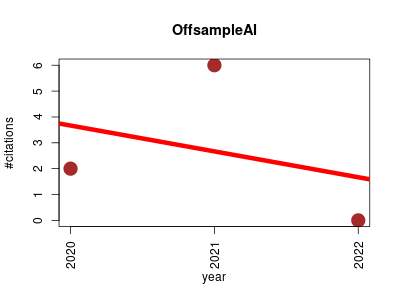 | 2 |
| maplet | Metabolomics Transcription factors and regulatory sites Workflows | An extensible R toolbox for modular and reproducible metabolomics pipelines. | -- | 8 | 0.0 | 8.0 | 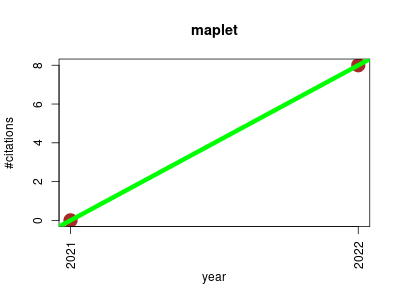 | 0 |
| Netsplitter | Biochemistry Metabolomics Molecular interactions, pathways and networks Systems biology | Mathematica software application to split biochemical networks into functional subnetworks. | 2011-02 | 7 | 0.6 | -0.1 | 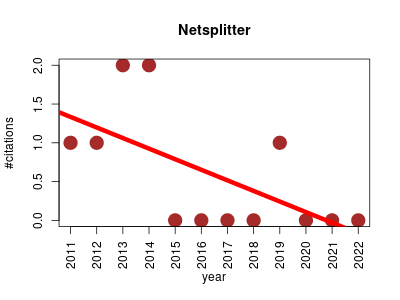 | 1 |
| BQuant | Data management Metabolomics NMR | R package using a probabilistic approach for fully automated database-based identification and quantification of metabolites in local regions of 1H Nuclear Magnetic Resonance (NMR) spectra. The approach is based on a linear mixed model, which accounts for technological characteristics of NMR spectra, and represents the spectra as mixtures of reference profiles from a database. | 2011-06 | 7 | 0.6 | 0.2 | 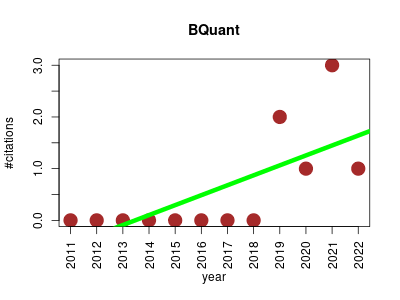 | 0 |
| PROM | Biology Genotype and phenotype Metabolomics Nucleic acid sites, features and motifs Systems biology | Algorithm which represents the successful integration of a genome scale transcriptional regulatory network with a biochemically detailed metabolic network, bridging two important classes of systems biology models that have rarely been combined quantitatively. | 2013-01 | 7 | 0.7 | -0.1 | 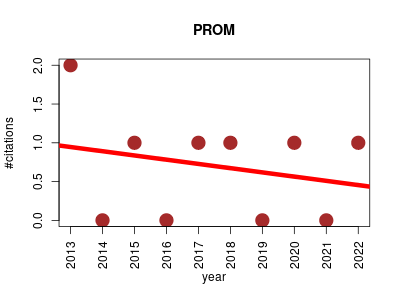 | 3 |
| AMBIENT | Metabolomics Molecular interactions, pathways and networks Systems biology | The Active Modules for Bipartite Networks is a Python module that uses simulated annealing to find areas of a metabolic network (modules) that have some consistent characteristic. It does not require predefined pathways and gives highly specific predictions of affected areas of metabolism. | 2013-03 | 7 | 0.7 | -0.2 | 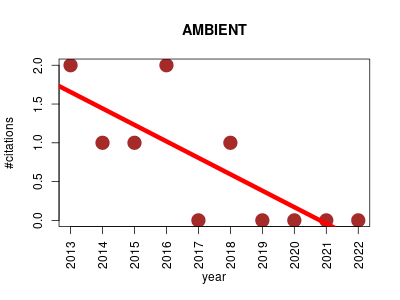 | 2 |
| MAIT | Laboratory techniques Metabolomics | This package contains functions to perform end-to-end statistical analysis of LC/MS Metabolomic Data. Special emphasis is put on peak annotation and in modular function design of the functions. | 2014-07 | 7 | 0.8 | 0.2 | 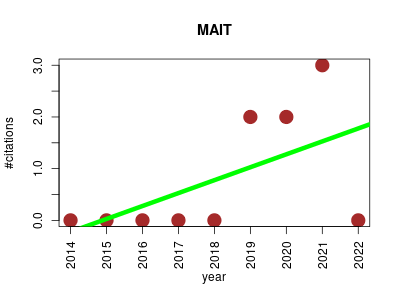 | 2 |
| MWASTools | Cheminformatics Lipids Metabolomics Systems biology | It provides a complete pipeline to perform metabolome-wide association studies. Key functionalities of the package include: quality control analysis of metabonomic data; MWAS using different association models (partial correlations; generalized linear models); model validation using non-parametric bootstrapping; visualization of MWAS results; NMR metabolite identification using STOCSY; and biological interpretation of MWAS results. | 2018-03 | 7 | 1.4 | 0.5 | 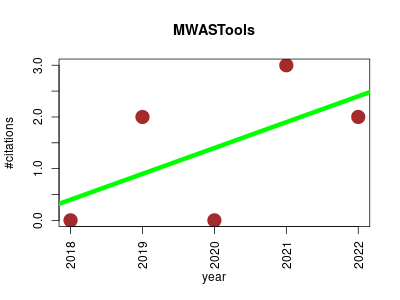 | 3 |
| pyQms | Metabolomics Proteomics | pyQms enables universal and accurate quantification of mass spectrometry data | 2017-10 | 7 | 1.2 | -0.3 | 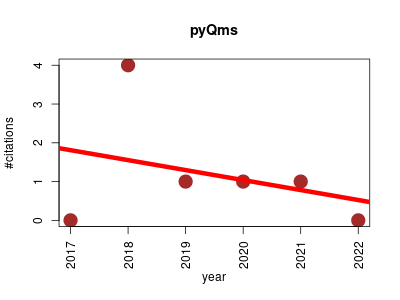 | 0 |
| pseudoQC | Metabolomics Proteomics Proteomics experiment | A Regression-Based Simulation Software for Correction and Normalization of Complex Metabolomics and Proteomics Datasets | Regression-Based Simulation Applications for Correction and Normalization of Complex Metabolomics and Proteomics Datasets | Wang, Shisheng, and Hao Yang. pseudoQC: A Regression‐Based Simulation Software for Correction and Normalization of Complex Metabolomics and Proteomics Datasets. Proteomics (2019): 1900264. (DOI: 10.1002/pmic.201900264) | 2019-10 | 7 | 1.8 | 0.7 | 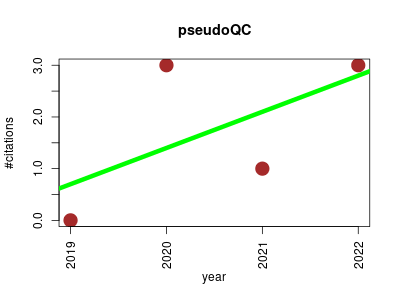 | 4 |
| IPA | Metabolomics Proteomics experiment | A Bayesian-Based Annotation Method for Metabolomic Profiles Integrating Biochemical Connections, Isotope Patterns, and Adduct Relationships | Integrated Probabilistic Annotation (IPA) - A Bayesian annotation method for LC/MS data integrating biochemical relations, isotope patterns and adduct formation | 2019-10 | 7 | 1.8 | 0.9 | 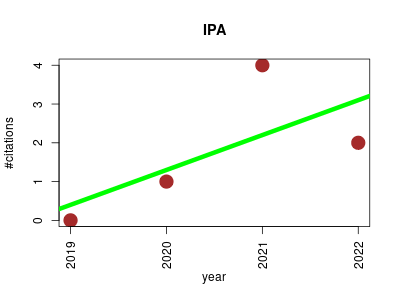 | 3 |
| LUCIDus | Biomarkers Genotype and phenotype Metabolomics | A Latent Unknown Clustering Integrating Multi-Omics Data (LUCID) with Phenotypic Traits | Latent Unknown Clustering with Integrated Data | An implementation for the LUCID method to jointly estimate latent unknown clusters/subgroups with integrated data. An EM algorithm is used to obtain the latent cluster assignment and model parameter estimates. Feature selection is achieved by applying the regularization method | 2020-02 | 7 | 2.3 | -1.5 | 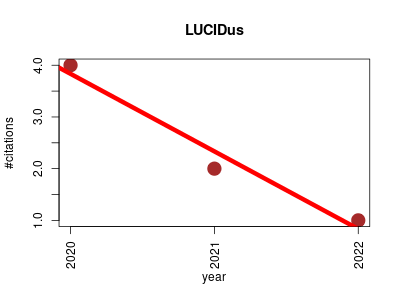 | 2 |
| Metandem | Metabolomics Proteomics Proteomics experiment Small molecules | An online software tool for mass spectrometry-based isobaric labeling metabolomics. | 2019-12 | 7 | 1.8 | -0.3 | 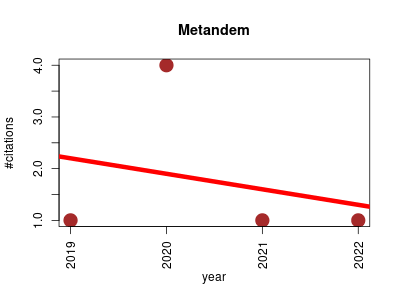 | 3 |
| MetWork | Compound libraries and screening Metabolomics Small molecules | Computer-Assisted Natural Products Anticipation | a web server for natural products anticipation | Currently running beta version 0.4.2 | Essence is a function of the relationship. Bachelard 1929 | MetWork used for the identification of bromotryptamine derivatives | 2019-09 | 7 | 1.8 | 0.7 | 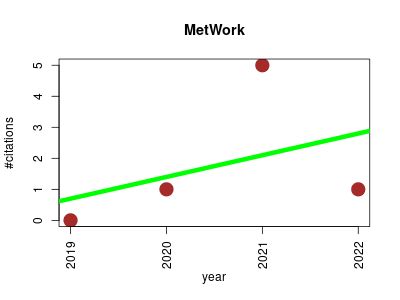 | 3 |
| TOXcms | Endocrinology and metabolism Medicinal chemistry Metabolomics Pharmacology Small molecules | Dose-Response Metabolomics to Understand Biochemical Mechanisms and Off-Target Drug Effects with the TOXcms Software. | 2020-01 | 7 | 2.3 | 0.5 | 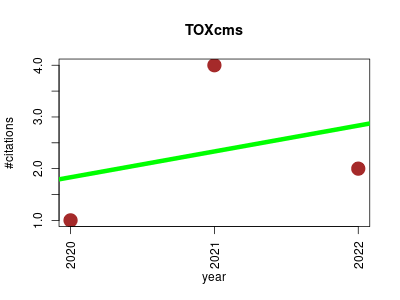 | 2 |
| apLCMS | Endocrinology and metabolism Metabolomics Proteomics experiment RNA-Seq Workflows | The R package apLCMS is designed for the processing of LC/MS based metabolomics data. It starts with a group of LC/MS files in the same folder, and generates a table with features in the rows and intensities in the columns. | 2020-12 | 7 | 2.3 | 3.0 | 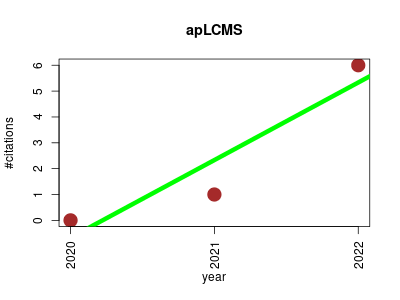 | 2 |
| MESSAR | Biochemistry Drug metabolism Metabolomics Proteomics experiment Small molecules | Automated recommendation of metabolite substructures from tandem mass spectra. | 2020-01 | 7 | 2.3 | 1.0 | 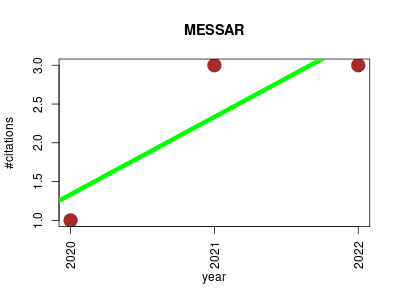 | 6 |
| LipidSig | Genotype and phenotype Lipids Machine learning Metabolomics Workflows | LipidSig is a web-based tool for lipidomic data analysis. LipidSig is the first web-based platform which integrates a comprehensive analysis for streamlined data mining of lipidomic datasets. The user-friendly interface provides five main functions, namely Profiling, Differential expression, Machine learning, Correlation and Network, for assessment of lipid effects on biological mechanisms. The five functions provide unique aspects to analyze the lipidome profiling data based on different characteristics including lipid class, chain length, unsaturation, hydroxyl group, and fatty acid composition., In summary, LipidSig enables users to perform intensive lipid analysis and create interactive plots with downloadable images and corresponding tables. | 2021-07 | 7 | 3.5 | 5.0 | 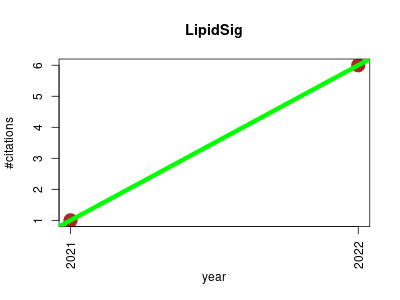 | 2 |
| PALS | Endocrinology and metabolism Metabolomics Molecular interactions, pathways and networks Proteomics experiment Small molecules | PALS (Pathway Activity Level Scoring) is a complete tool that performs database queries of pathways, decomposes activity levels in pathways via the PLAGE method, as well as presents the results in a user-friendly manner. The results are found to be more robust to noise and missing peaks compared to the alternatives (ORA, GSEA). This is particularly important for metabolomics peak data, where noise and missing peaks are prevalent. | 2021-02 | 7 | 3.5 | -3.0 | 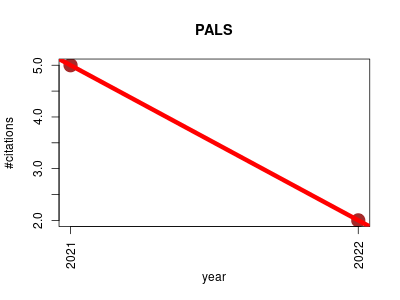 | 3 |
| MetCirc | Metabolomics Molecular interactions, pathways and networks Systems biology | This tool comprises a workflow to interactively explore metabolomics data by creating MSP or calculating m/z values and the similarity between precursors. | 2017-08 | 6 | 1.0 | -0.0 | 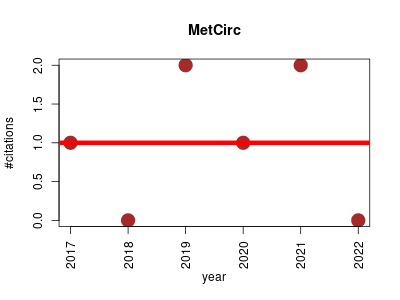 | 4 |
| MetaFIND | Metabolomics NMR | Java-based feature analysis tool that enables real-time, interactive correlation analysis of feature sets, discovery of addtional related features and analysis of related metabolites. It is designed specifically to aid feature analysis within NMR metabolomics data. | 2008-11 | 6 | 0.4 | -0.0 | 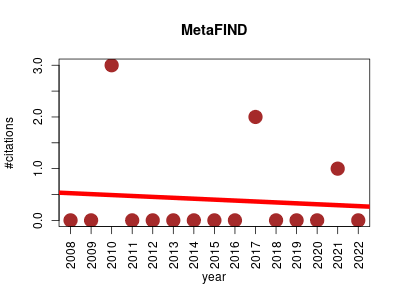 | 2 |
| mmnet | Metabolomics Microbiology Molecular interactions, pathways and networks Systems biology | This package gives the implementations microbiome metabolic network constructing and analyzing. It introduces a unique metagenomic systems biology approach, mapping metagenomic data to the KEGG global metabolic pathway and constructing a systems-level network. The system-level network and the next topological analysis will be of great help to analysis the various functional properties, including regulation and metabolic functionality of the metagenome. | 2015-01 | 6 | 0.8 | -0.1 | 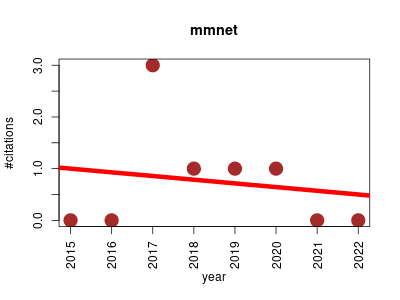 | 1 |
| compareMS2 | Metabolomics Phylogeny Proteomics Proteomics experiment | A simple tool developed to compare, globally, all MS/MS spectra between two datasets (in Mascot Generic Format or MGF). | 2012-04 | 6 | 0.5 | 0.1 | 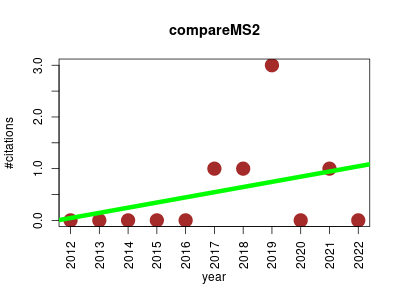 | 0 |
| OpenMZxy | Biological imaging Lipids Metabolomics Proteomics Proteomics experiment | Open mass spectrometry imaging control software. | 2014-05 | 6 | 0.7 | 0.1 | 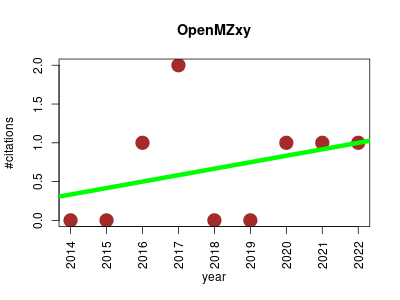 | 1 |
| SIMAT | Data visualisation Metabolomics | This package provides a pipeline for analysis of GC-MS data acquired in selected ion monitoring (SIM) mode. The tool also provides a guidance in choosing appropriate fragments for the targets of interest by using an optimization algorithm. This is done by considering overlapping peaks from a provided library by the user. | 2015-08 | 6 | 0.8 | -0.3 | 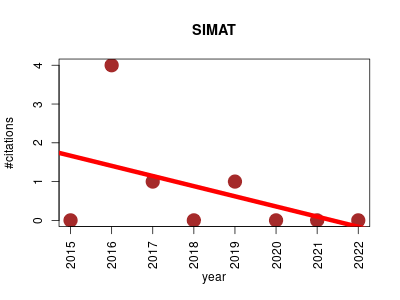 | 2 |
| iMet-Q | Metabolomics | iMet-Q (intelligent Metabolomic Quantitation) is a software that quantifies metabolites in full-scan liquid chromatography-mass spectrometry (LC-MS) data. | 2016-01 | 6 | 0.9 | 0.2 | 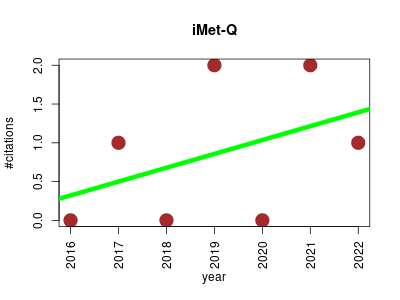 | 3 |
| WebSpecmine | Bioinformatics Data mining Data visualisation Database management Machine learning Metabolomics | WebSpecmine is a web-based application designed to perform the analysis of metabolomics data based on spectroscopic and chromatographic techniques (NMR, Infrared, UV-visible, and Raman, and LC/GC-MS)) and compound concentrations. | 2019-10 | 6 | 1.5 | 0.4 | 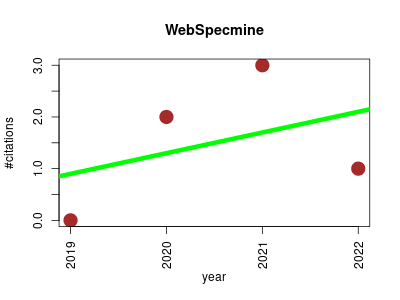 | 3 |
| Fusion | Bioinformatics Data integration and warehousing Data visualisation Gene expression Gene regulation Metabolomics Protein expression Proteomics Transcriptomics | Web application for the integrative analysis of omics data providing a collection of new and established tools and visualization methods to support researchers in exploring multi-level omics data. The software focuses on data-rich high-throughout experiments from transcriptomics, proteomics and/or metabolomics, and offers convenient functionality for a) omics data manipulation, b) data analysis, c) cluster analysis, d) data visualization. | 2016-12 | 6 | 0.9 | 0.4 | 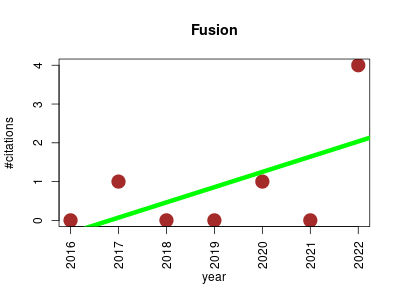 | 1 |
| Midcor | Fluxomics Metabolomics | R-program supporting a step of workflow of fluxomic analysis of artificial 13C labeling of metabolites. It designed to correct raw mass spectra (MS) of 13C-labeled metabolites of interest for natural isotopes occurrence. The raw mass spectra are supposed to be extracted from MS recording by Ramid. | 2017-02 | 6 | 1.0 | 0.6 | 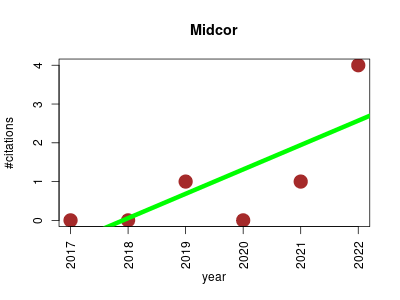 | 2 |
| iSMU | Endocrinology and metabolism Metabolomics Microbial ecology Molecular interactions, pathways and networks Small molecules | Metabolic Modeling of Streptococcus mutans Reveals Complex Nutrient Requirements of an Oral Pathogen. | 2019-01 | 6 | 1.5 | 0.4 | 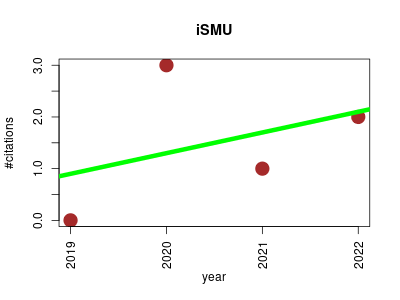 | 0 |
| LipMat | Lipids Metabolomics NMR Proteomics Proteomics experiment | Fast and Quantitative Phospholipidomic Analysis of SH-SY5Y Neuroblastoma Cell Cultures Using Liquid Chromatography-Tandem Mass Spectrometry and 31P Nuclear Magnetic Resonance. | 2019-12 | 6 | 1.5 | 0.8 | 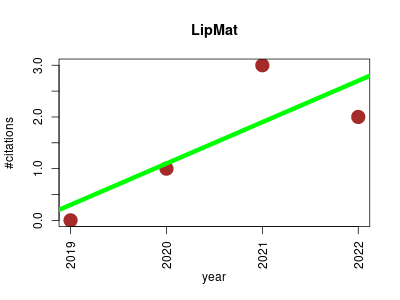 | 0 |
| VOCCluster | Endocrinology and metabolism Metabolomics Proteomics Proteomics experiment Small molecules | Untargeted Metabolomics Feature Clustering Approach for Clinical Breath Gas Chromatography. | 2020-02 | 6 | 2.0 | 0.0 |  | 3 |
| WiPP | Machine learning Metabolomics Workflows | Workflow for Improved Peak Picking for Gas Chromatography-Mass Spectrometry (GC-MS) Data | large scale GC-MS data preprocessing workflow | WiPP - A Workflow for improved Peak Picking - Quick Start Guide | WiPP is an open source large scale GC-MS data preprocessing workflow built in Python 3 that uses machine learning to optimise, automate and combine the peak detection process of commonly used peak picking algorithms | 2019-09 | 6 | 1.5 | 0.6 |  | 2 |
| AlpsNMR | Endocrinology and metabolism Metabolomics NMR Small molecules Workflows | AlpsNMR is an R package that can load Bruker and JDX samples as well as preprocess them. | 2020-05 | 6 | 2.0 | 0.5 |  | 4 |
| Planet Microbe | Metabolomics Metagenomics Metatranscriptomics Microbiology Ontology and terminology | Planet Microbe is a platform for marine microbiology to discover and analyze interconnected omics and environmental data. | 2021-01 | 6 | 3.0 | 0.0 |  | 0 |
| PUMA | Endocrinology and metabolism Machine learning Metabolomics Molecular interactions, pathways and networks Small molecules | Pathway-Activity Likelihood Analysis and Metabolite Annotation for Untargeted Metabolomics Using Probabilistic Modeling. | 2020-05 | 6 | 2.0 | 0.5 | 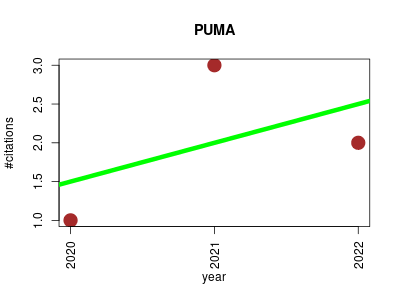 | 3 |
| CROP | Bioinformatics Metabolomics Proteomics experiment | CROP (Correlation-based Removal Of multiPlicities) is a visual post-processing tool that removes redundant features from untargeted metabolomic data sets. It is based on a grouping of highly correlated features within a defined retention time window avoiding the condition of specific m/z difference making it a second-tier strategy for multiplicities reduction. Graphical representation of correlation network for better understanding of the clusters composition and parameter tuning is provided. | 2020-05 | 6 | 2.0 | 0.5 | 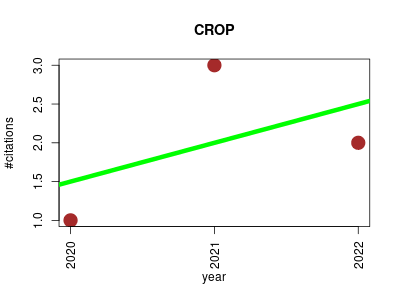 | 1 |
| MetaboKit | Compound libraries and screening Lipids Metabolomics Proteomics experiment Small molecules | A comprehensive data extraction tool for untargeted metabolomics. | 2020-10 | 6 | 2.0 | -1.0 | 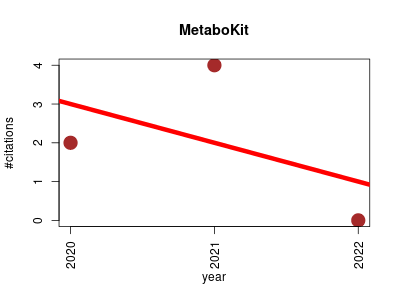 | 0 |
| struct | Machine learning Metabolomics Proteomics experiment Statistics and probability Workflows | an R/bioconductor-based framework for standardised metabolomics data analysis and beyond. | 2020-12 | 6 | 2.0 | 2.0 | 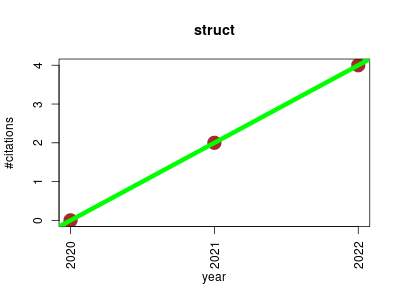 | 1 |
| SYNERGxDB | Biomarkers Drug discovery Metabolomics Oncology Pharmacogenomics | an integrative pharmacogenomic portal to identify synergistic drug combinations for precision oncology. | 2020-01 | 6 | 2.0 | 0.0 | 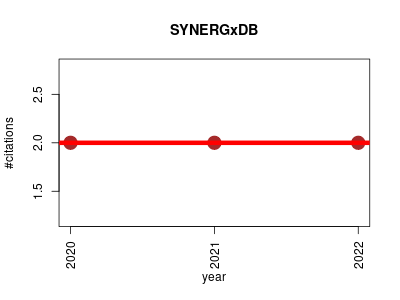 | 1 |
| Escher-Trace | Endocrinology and metabolism Metabolomics Molecular interactions, pathways and networks Proteomics experiment Small molecules | Escher-Trace is an interactive pathway visualization tool for on-the-fly stable isotope tracing analysis. | 2020-07 | 6 | 2.0 | 1.5 | 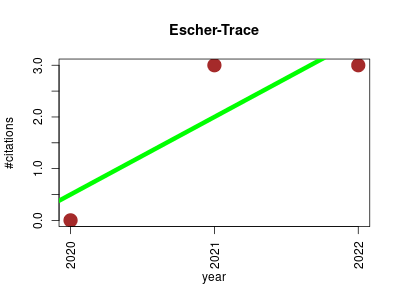 | 1 |
| XenoNet | Drug metabolism Endocrinology and metabolism Metabolomics Molecular interactions, pathways and networks Small molecules | Inference and Likelihood of Intermediate Metabolite Formation. | 2020-07 | 6 | 2.0 | -0.0 | 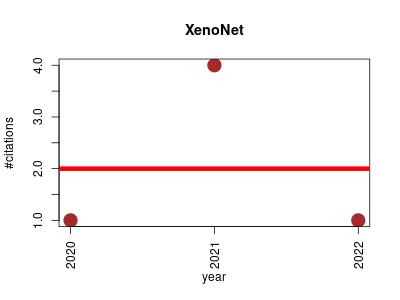 | 1 |
| gutSMASH | Endocrinology and metabolism Metabolomics Microbial ecology Molecular interactions, pathways and networks Small molecules | gutSMASH web server is an automated identification of primary metabolic gene clusters from the gut microbiota. It is an approach to functionally profile the human microbiome for specialized primary metabolic gene clusters. | -- | 6 | 0.0 | 4.0 | 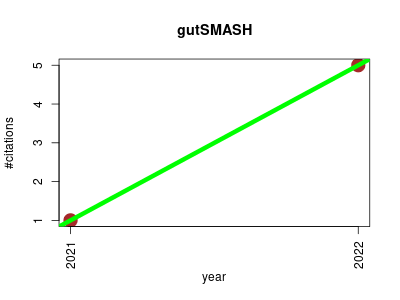 | 1 |
| iNetModels | Metabolomics Metagenomics Molecular interactions, pathways and networks Oncology Proteomics | iNetModels is an interactive visualization and database of multi-omics data. | 2021-07 | 6 | 3.0 | 0.0 |  | 0 |
| peakPantheR | Endocrinology and metabolism Metabolomics Proteomics experiment Small molecules Workflows | R package for large-scale targeted extraction and integration of annotated metabolic features in LC-MS profiling datasets. | -- | 6 | 0.0 | 6.0 |  | 0 |
| PlantMetabolomics | Endocrinology and metabolism Metabolomics Molecular interactions, pathways and networks Small molecules | An NSF-funded multi-institutional project developing metabolomics as a functional genomics tool for elucidating the functions of Arabidopsis genes. | 2012-01 | 5 | 0.5 | -0.1 |  | 0 |
| MetAssimulo | Metabolomics Molecular medicine NMR | MATLAB-based package which simulates 1H-NMR spectra of complex mixtures such as metabolic profiles. Drawing data from a metabolite standard spectral database in conjunction with concentration information input by the user or constructed automatically from the Human Metabolome Database. | 2010-10 | 5 | 0.4 | -0.0 |  | 1 |
| Ambit-SMIRKS | Cheminformatics Enzymes Metabolomics Protein structure analysis | Software module for reaction representation, reaction search and structure transformation. | 2018-12 | 5 | 1.0 | 0.3 | 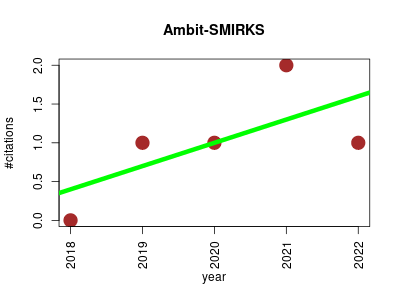 | 1 |
| yamss | Data visualisation Metabolomics | Tools to analyze and visualize high-throughput metabolomics data aquired using chromatography-mass spectrometry. These tools preprocess data in a way that enables reliable and powerful differential analysis. | 2017-03 | 5 | 0.8 | -0.1 | 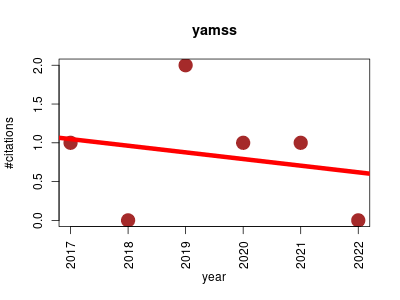 | 4 |
| Goslin | Bioinformatics Cheminformatics Computational chemistry Lipids Metabolomics Ontology and terminology | Goslin is the Grammar of succinct lipid nomenclature project. It defines multiple grammers compatible with ANTLRv4 for different sources of shorthand lipid nomenclature. This allows to generate parsers based on the defined grammars, which provide immediate feedback whether a processed lipid shorthand notation string is compliant with a particular grammar, or not. | 2020-08 | 5 | 1.7 | 2.0 | 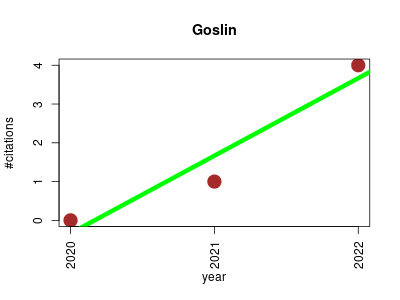 | 2 |
| MetGem | Metabolomics Molecular interactions, pathways and networks Proteomics experiment | Generation of a Molecular Network from Electron Ionization Mass Spectrometry Data by Combining MZmine2 and MetGem Software | Calculation and visualization of molecular networks based on t-SNE algorithm - metgem/metgem | MetGem is an open-source software for tandem mass-spectrometry data visualization. Its key features are standalone molecular networking and t-SNE based projections | 2019-09 | 5 | 1.2 | 0.7 | 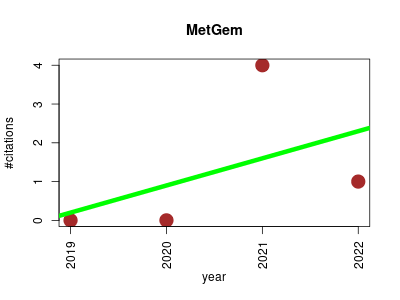 | 1 |
| moiety_modeling | Endocrinology and metabolism Metabolomics NMR Proteomics experiment Small molecules | Moiety modeling framework for deriving moiety abundances from mass spectrometry measured isotopologues. | 2019-10 | 5 | 1.2 | 0.1 | 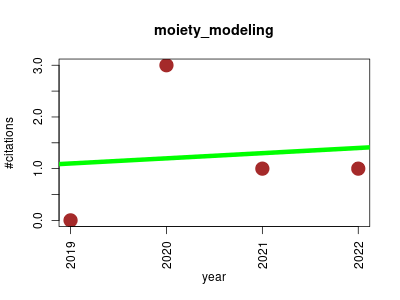 | 0 |
| FUMOSO | Cell biology Endocrinology and metabolism Metabolomics Oncology Pharmacology | Fuzzy modeling and global optimization to predict novel therapeutic targets in cancer cells. | 2020-04 | 5 | 1.7 | 1.5 | 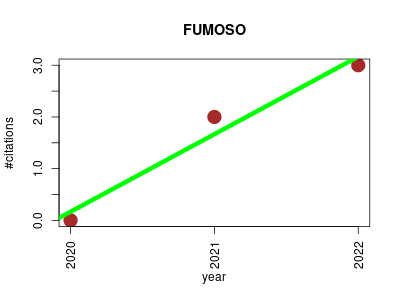 | 0 |
| Pathmodel | Metabolomics Systems biology | PathModel is a prototype to infer new biochemical reactions and new metabolite structures. | 2020-02 | 5 | 1.7 | 1.0 | 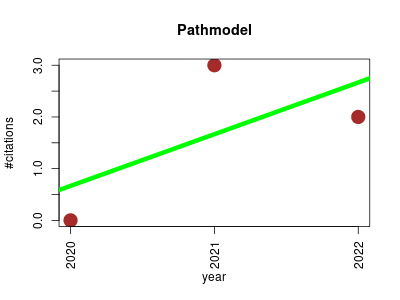 | 2 |
| CycloBranch | Imaging Metabolomics Proteomics Proteomics experiment Small molecules | CycloBranch is an open and platform-free software that provide the de novo generation of molecular formulas of unknown compounds in both liquid chromatography/mass spectrometry and mass spectrometry imaging datafiles. | 2020-05 | 5 | 1.7 | -1.5 | 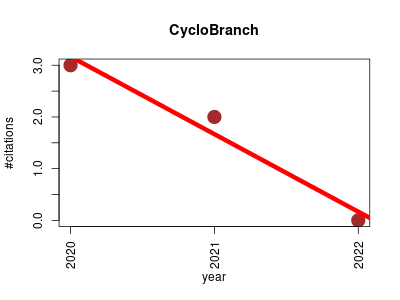 | 0 |
| NetMet | Endocrinology and metabolism Metabolomics Microbial ecology Nutritional science Small molecules | A Network-Based Tool for Predicting Metabolic Capacities of Microbial Species and their Interactions. | 2020-06 | 5 | 1.7 | 0.5 | 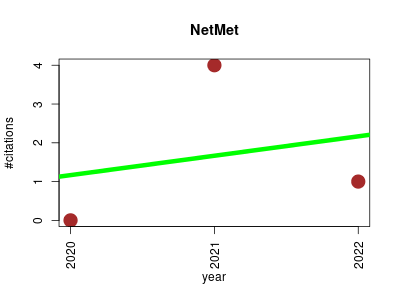 | 2 |
| Fluxer | Endocrinology and metabolism Fluxomics Metabolomics Molecular interactions, pathways and networks Small molecules | Fluxer is a web application for the computation and interactive visualization of flux graphs from genome-scale metabolic models. Flux Balance Analysis is used to calculate the flux of the metabolic network, which can then be visualized with spanning trees, k-shortest paths, and complete graphs | 2020-01 | 5 | 1.7 | 0.5 | 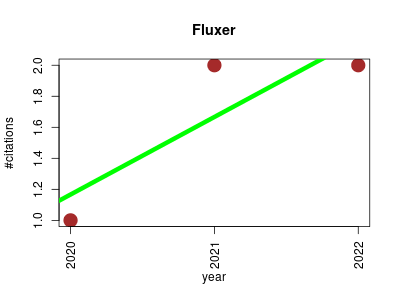 | 4 |
| SeMPI | Machine learning Metabolomics Molecular modelling Small molecules | Web Server for PKS and NRPS Predictions Combined with Metabolite Screening in Natural Product Databases. | 2020-01 | 5 | 1.7 | -0.0 | 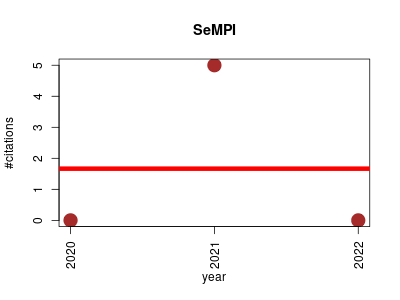 | 2 |
| AutoCCS | Metabolomics Proteomics experiment Workflows | Automated Collision Cross Section calculation software for ion mobility-mass spectrometry | -- | 5 | 0.0 | 5.0 | 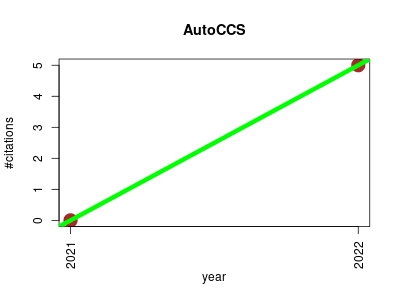 | 1 |
| novoSpaRc | Cell biology Gene expression Gene transcripts Metabolomics RNA-Seq | novoSpaRc predicts locations of single cells in space by solely using single-cell RNA sequencing data. An existing reference database of marker genes is not required, but significantly enhances performance if available. | 2021-09 | 5 | 2.5 | 5.0 |  | 1 |
| timeOmics | Gene transcripts Metabolomics Microbial ecology Molecular biology Small molecules | timeOmics is a generic data-driven framework to integrate multi-Omics longitudinal data measured on the same biological samples and select key temporal features with strong associations within the same sample group. | -- | 5 | 0.0 | 5.0 |  | 1 |
| FROMP | Mapping Metabolomics Molecular interactions, pathways and networks Systems biology | Software for mapping and visualizing enzyme annotations onto the Kyoto Encyclopedia of Genes and Genomes (KEGG) metabolic pathways or custom-made pathways and comparing the samples in terms of their Pathway Completeness Scores, their relative Activity Scores or enzyme enrichment odds ratios. | 2013-03 | 4 | 0.4 | -0.1 |  | 1 |
| sampleDrift | Data quality management Metabolomics NMR | A package to explore the effects of pre-centrifugation storage on plasma sample quality. | 2017-11 | 4 | 0.7 | 0.1 |  | 2 |
| Miso | Metabolomics Small molecules | R Package for Multiple Isotope Labeling Assisted Metabolomics Data Analysis. | 2019-09 | 4 | 1.0 | -0.4 |  | 2 |
| ARISTO | Metabolomics Ontology and terminology Small molecules | Web tool which provides information regarding the chemical nature/ontology of the compound underlying an input electron ionization mass spectrum. | 2011-07 | 4 | 0.3 | -0.0 | 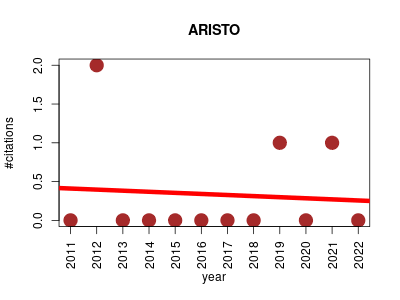 | 1 |
| CluMSID | Metabolomics Proteomics experiment Small molecules | R package for similarity-based clustering of tandem mass spectra to aid feature annotation in metabolomics. | 2019-09 | 4 | 1.0 | 0.2 | 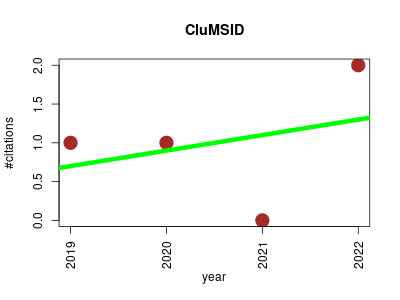 | 1 |
| ADAPTIVE | Machine learning Metabolomics Small molecules | leArning DAta-dePendenT, concIse molecular VEctors for fast, accurate metabolite identification from tandem mass spectra | Learning molecular vectors for metabolites from spectra-structure pairs] | We propose ADAPTIVE, which has two parts: learning two mappings 1) from structures to molecular vectors and 2) from spectra to molecular vectors | 2019-07 | 4 | 1.0 | 0.6 | 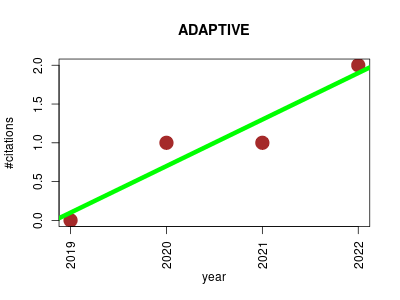 | 2 |
| GRaMM | Metabolomics Microbial ecology Pathology Physiology Small molecules | Strategy for Intercorrelation Identification between Metabolome and Microbiome. | 2019-01 | 4 | 1.0 | 0.2 | 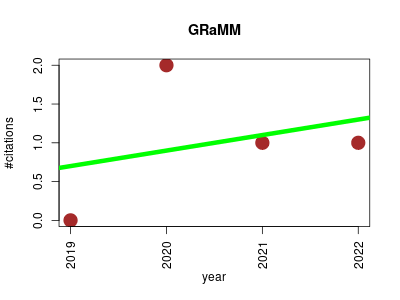 | 1 |
| IonSpattern | Imaging Metabolomics Proteomics experiment | Unsupervised segmentation of mass spectrometric ion images characterizes morphology of tissues | Spatial Dirichlet Gaussian Mixture Model for single ion images | run W_matrix.R GMM.R k_DGMM.R S_DGMM.R first | run CpG_Sal_mouse brain.R to get results of CpG and Sal preconditioned mouse brain data | 2019-07 | 4 | 1.0 | 0.8 | 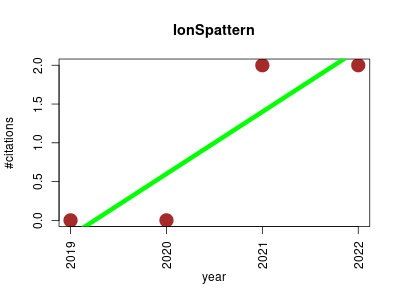 | 0 |
| jmztab-m | Data submission, annotation and curation Metabolomics Ontology and terminology | Reference implementation for mzTab 2.0 for metabolomics | 2019-10 | 4 | 1.0 | 0.4 | 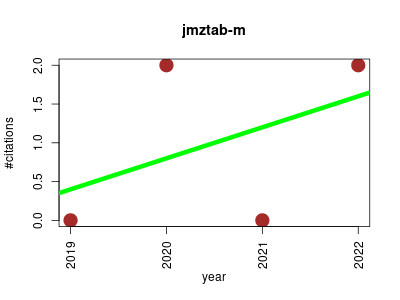 | 3 |
| NMR | Metabolomics NMR RNA | Single-sample quantification of RNA, proteins and metabolites for biomolecular network analysis | In Vitro Transcription NMR. Protocol, code and examples for the co-transcriptional RNA folding network reconstruction | The protocol describes how to setup and analyse observation of a co-transcriptional RNA folding network by Systems NMR approach. While most experimental approaches can monitor only a single molecule class or reaction type at a time, Systems NMR permits single-sample dynamic quantification of entire “heterotypic” networks – involving different reaction and molecule types. It thus provides a deeper systems-level understanding of biological network dynamics by combining the dynamic resolution of biochemical assays and the multiplexing ability of “omics” | 2019-08 | 4 | 1.0 | 0.2 | 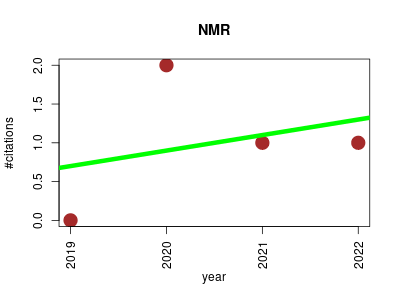 | 1 |
| Argonaut | Lipids Metabolomics Proteomics Proteomics experiment Sequence analysis | Argonaut is a web-based utility which facilitates collaborative exploration of Condition/Control-type mass spectrometry experiments. Spreadsheets of quantitative values can be uploaded and organized in a way that best describes your specific experimental setup. Brief statistical analysis is conducted on uploaded data to calculate the significance of molecular perturbations for each condition. These data can then be queried on-the-fly to conduct commonly conducted analyses inside the data portal, including volcano plots, GO enrichment, linear correlations, and others. | 2020-10 | 4 | 1.3 | 1.0 |  | 0 |
| KLIC | ChIP-on-chip Epigenomics Gene expression Machine learning Metabolomics | Multiple kernel learning for integrative consensus clustering of omic datasets. | 2020-09 | 4 | 1.3 | 1.0 | 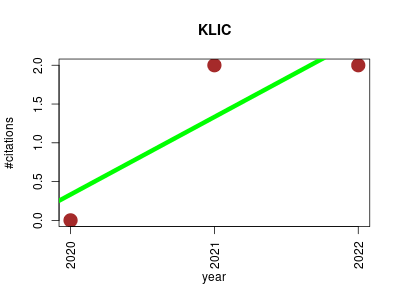 | 2 |
| SmartPeak | Fluxomics Lipids Metabolomics Proteomics experiment Workflows | SmartPeak Automates Targeted and Quantitative Metabolomics Data Processing. | 2020-12 | 4 | 1.3 | 0.5 | 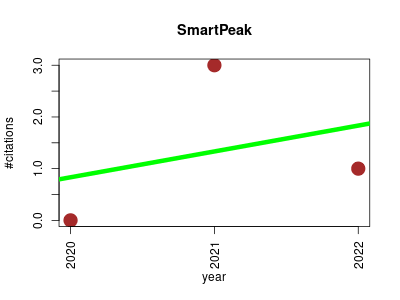 | 2 |
| SMfinder | Lipids Metabolomics Molecular biology Proteomics experiment Small molecules | Small Molecules Finder for Metabolomics and Lipidomics Analysis. | 2020-07 | 4 | 1.3 | -0.0 |  | 2 |
| MetaboMAPS | Endocrinology and metabolism Metabolomics Molecular interactions, pathways and networks Proteomics Small molecules | Pathway sharing and multi-omics data visualization in metabolic context. | -- | 4 | 0.0 | -0.5 | 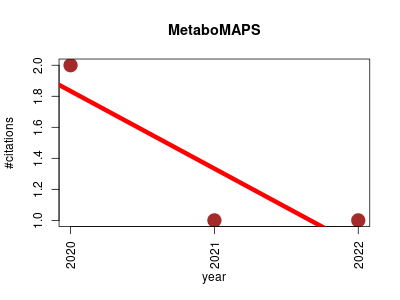 | 0 |
| NECom | Biotechnology Endocrinology and metabolism Metabolomics Model organisms Small molecules | Predicting Nash equilibria for microbial metabolic interactions. | 2020-12 | 4 | 1.3 | -0.0 | 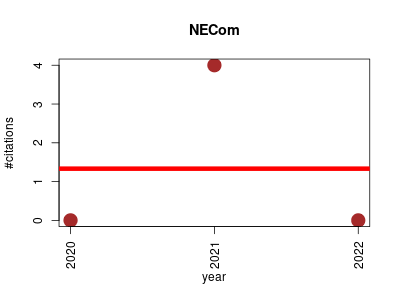 | 0 |
| DynLib | Endocrinology and metabolism Metabolomics Molecular interactions, pathways and networks NMR Proteomics experiment | DynLib is spectral database containing spectras from LC-MS runs for maize organs in order to investigate the metabolome. | -- | 4 | 0.0 | 0.0 | 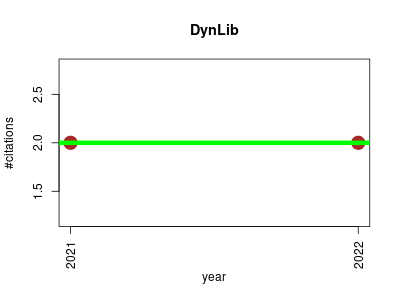 | 1 |
| LipidSuite | Endocrinology and metabolism Lipids Metabolomics Proteomics experiment Workflows | LipidSuite is an interactive web server for lipidomics differential and enrichment analysis. | 2021-07 | 4 | 2.0 | 2.0 | 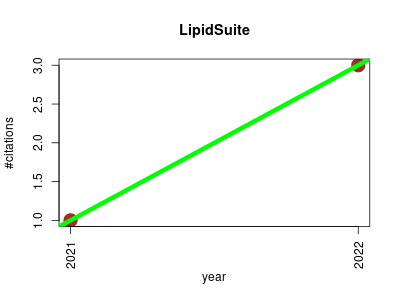 | 1 |
| omeClust | Mapping Metabolomics Metagenomics Pathology | omeClust is a clustering method that detects clusters of features using omics data and scores metadata (resolution score) based on their influences in clustering. The similarity of features within each cluster can be different (different resolution). Resolution of similarity score takes to account not only similarity between measurements and also the structure in a hierarchical structure of data and number of features which group together. | -- | 4 | 0.0 | 0.0 | 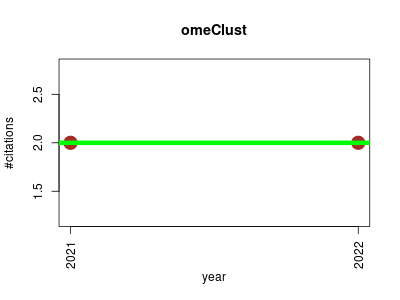 | 0 |
| PSC-db | Drug metabolism Metabolomics Molecular modelling Plant biology Small molecules | A structured and searchable 3D-Database for Plant Secondary Compounds. | 2021-02 | 4 | 2.0 | 0.0 | 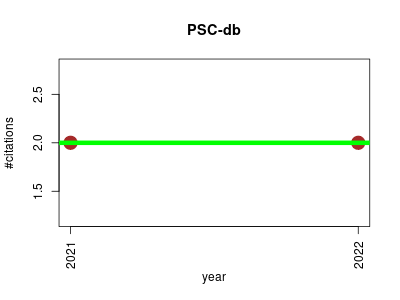 | 3 |
| ISFrag | Endocrinology and metabolism Metabolomics Molecular interactions, pathways and networks Proteomics experiment Small molecules | De Novo Recognition of In-Source Fragments for Liquid Chromatography-Mass Spectrometry Data. | 2021-07 | 4 | 2.0 | 4.0 | 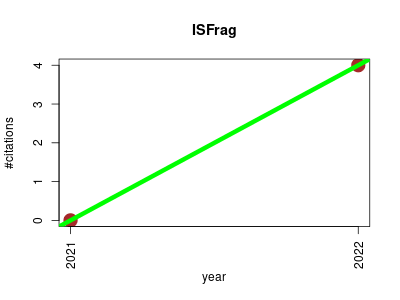 | 1 |
| MS2Compound | Bioinformatics Biotherapeutics Metabolomics | MS2Compound is a user-friendly compound identification tool for LC-MS/MS-based metabolomics data. | 2021-06 | 4 | 2.0 | 2.0 | 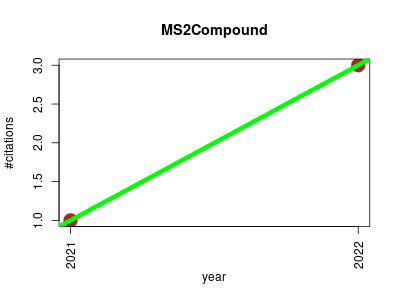 | 0 |
| PIXiE | Genomics Metabolomics Proteomics experiment | Algorithm for automated ion mobility arrival time extraction and collision cross section calculation using global data association. | 2017-09 | 4 | 0.7 | -0.1 | 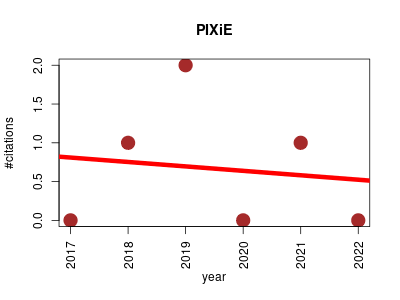 | 1 |
| NP-MRD | Biochemistry Medicinal chemistry Metabolomics NMR Small molecules | The NP-MRD is a freely available cloud-based, user-friendly, FAIR electronic database. NP-MRD accepts NMR data and associated metadata from newly undertaken NP studies. | 2022-01 | 4 | 4.0 | 0.0 | 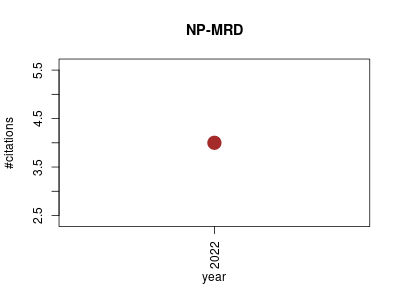 | 1 |
| LINT-web | Endocrinology and metabolism Lipids Metabolomics Proteomics Transcriptomics | A lipidomic data processing website aims to provide users a friendly pipeline for statistical analyses and lipidomic data mining. | 2021-09 | 4 | 2.0 | 4.0 | 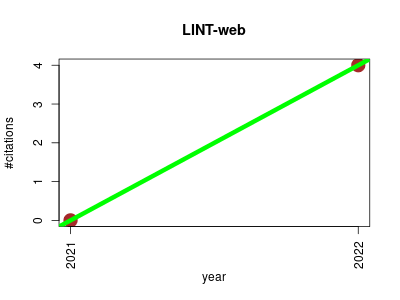 | 0 |
| IMSDB | Data management Data mining Metabolomics | Comprehensive database application and analysis platform that is capable of combining metabolic data, like IMS chromatograms, and heterogeneous biomedical data, like patient records, into a centralized data structure. | 2013-01 | 3 | 0.3 | -0.0 |  | 0 |
| INDEED | Biomarkers Metabolomics Proteomics | R package for network based differential expression analysis. | 2019-01 | 3 | 0.8 | 0.1 | 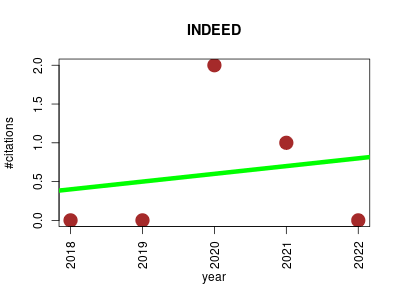 | 2 |
| GGMs | Metabolomics Molecular interactions, pathways and networks Statistics and probability | Exact hypothesis testing for shrinkage based Gaussian Graphical Models. | 2019-12 | 3 | 0.8 | -0.5 | 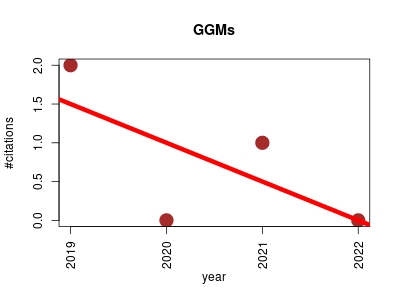 | 1 |
| DynaStI | Compound libraries and screening Lipids Metabolomics | Dynamic Retention Time Database for Steroidomics. | 2019-05 | 3 | 0.8 | 0.1 | 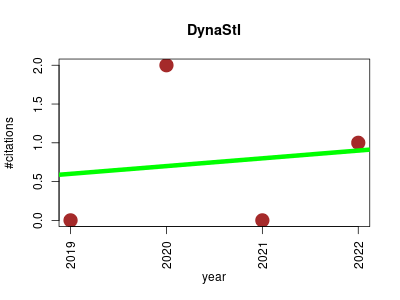 | 2 |
| KBCommons | Epigenomics Functional, regulatory and non-coding RNA Metabolomics Phenomics Proteomics | a universal framework for multi-omics data integration and biological discoveries. | 2019-12 | 3 | 0.8 | 0.1 | 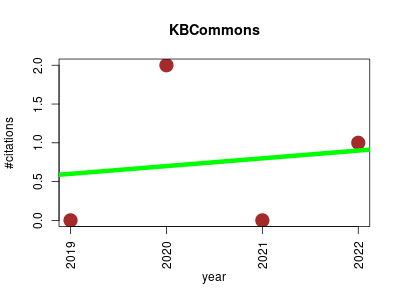 | 2 |
| MetaboList | Metabolomics Proteomics experiment Small molecules | A Flexible Tool for Metabolite Annotation from High-Resolution Data-Independent Acquisition Mass Spectrometry Analysis | Annotation of Metabolites from Liquid Chromatography-Mass Spectrometry Data | Automatic metabolite annotation from Liquid Chromatography-Mass Spectrometry (LC-MS and LC-MS/MS DIA) analysis | 2019-09 | 3 | 0.8 | 0.1 | 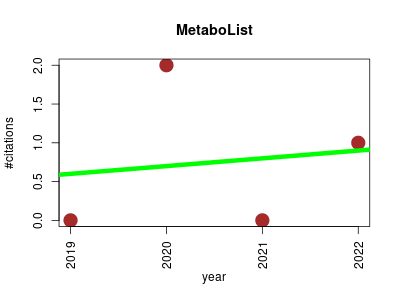 | 1 |
| MetumpX | Metabolomics Proteomics experiment Workflows | A Metabolomics Support Package for Untargeted Mass Spectrometry. | 2020-03 | 3 | 1.0 | 0.0 | 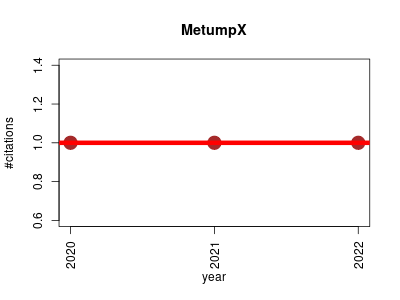 | 3 |
| PySpacell | Cell biology Genotype and phenotype Imaging Metabolomics Proteomics experiment | PySpacell is a Python Package for Spatial Analysis of Cell Images. | 2020-03 | 3 | 1.0 | 1.5 | 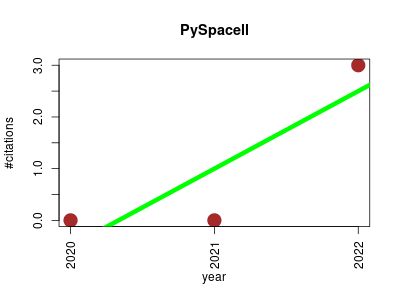 | 1 |
| Metabolite-Investigator | Endocrinology and metabolism Metabolomics Small molecules Workflows | An integrated userfriendly workflow for metabolomics multi-study analysis. | 2021-08 | 3 | 1.5 | 1.0 | 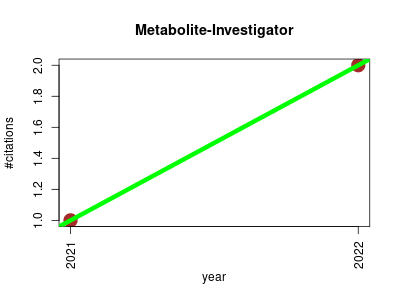 | 1 |
| DeepAE | Cytometry Gene expression Metabolomics Pathology RNA-Seq | DeepAE is deep learning framework to identify the key dimensions from high-dimensional gene expression profiles. DeepAE is composed of an input layer, seven hidden layers, and output layer to form the encoder and decoder phases that are corresponded to the compression and decompression. | 2021-01 | 3 | 1.5 | 1.0 | 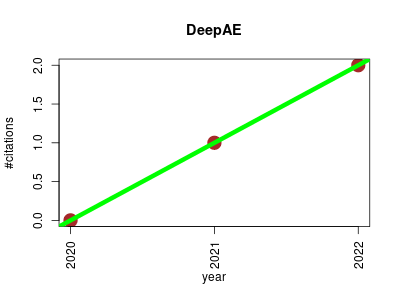 | 1 |
| DIMEDR | Metabolomics Proteomics Proteomics experiment Small molecules | DIMEDR (Disparate Metabolomics Data Reassembler) is a algorithm for agglomerating incongruent LC-MS metabolomics datasets. (DIMEDR) attempts to bridge the inconsistencies between incongruent LC-MS metabolomics datasets of the same biological sample type. A single “primary” dataset is postprocessed via traditional means of peak identification, alignment, and grouping. DIMEDR utilizes this primary dataset as a progenitor template by which data from subsequent disparate datasets are reassembled and integrated into a unified framework that maximizes spectral feature similarity across all samples. This is accomplished by a novel procedure for universal retention time correction and comparison via identification of ubiquitous features in the initial primary dataset, which are subsequently utilized as endogenous internal standards during integration. | 2020-04 | 3 | 1.0 | 0.0 | 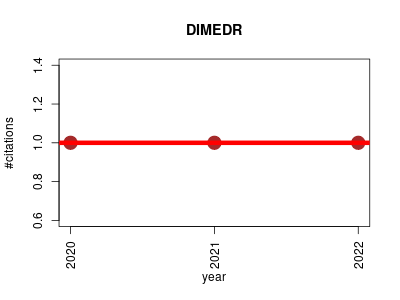 | 2 |
| FastMM | Endocrinology and metabolism Metabolomics Molecular interactions, pathways and networks Oncology Small molecules | FastMM is an efficient toolbox for personalized constraint-based metabolic modeling. In FastMM, most of the time-cost functions of metabolic modeling are written by C. These functions including: flux variability analysis, genome-wide single gene knockout analysis, genome-wide double gene knockout analysis, genome-wide single metabolite knockout analysis, genome-wide double metabolite knockout analysis, and MCMC sampling. | 2020-02 | 3 | 1.0 | -0.5 | 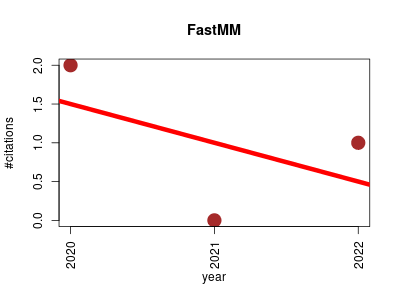 | 1 |
| venoms | Literature and language Metabolomics Proteomics experiment Small molecules Taxonomy | Website for the Low Molecular Mass Compounds in Spider Venoms. | 2020-08 | 3 | 1.0 | 1.5 | 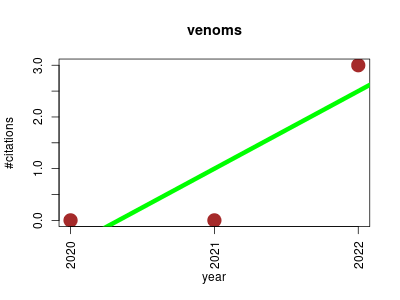 | 0 |
| MSI-R | Imaging Metabolomics Proteomics experiment Transcription factors and regulatory sites | Adaptive Pixel Mass Recalibration for Mass Spectrometry Imaging Based on Locally Endogenous Biological Signals. | 2021-03 | 3 | 1.5 | 3.0 |  | 1 |
| SistematX | Drug discovery Metabolomics Pharmacology Plant biology Small molecules | The SistematX Web Portal of Natural Products is database of secondary metabolites. | 2021-06 | 3 | 1.5 | 3.0 |  | 2 |
| SteroidXtract | Data mining Endocrinology and metabolism Metabolomics Proteomics experiment Small molecules | SteroidXtract is a deep-learning based bioinformatics tool for steroid MS2 extraction from untargeted metabolomics data. | 2021-04 | 3 | 1.5 | -1.0 |  | 0 |
| EVA | Endocrinology and metabolism Machine learning Metabolomics Proteomics experiment | EVA is a deep-learning-based program that automatically evaluates the fidelity of metabolic features generated from LC-MS experiments | -- | 3 | 0.0 | 3.0 |  | 0 |
| massPix | Imaging Metabolomics Proteomics experiment Statistics and probability | Processes high resolution mass spectrometry imaging data, performs multivariate statistics (PCA, clustering) and lipid identification. | 2017-11 | 3 | 0.5 | 0.0 | 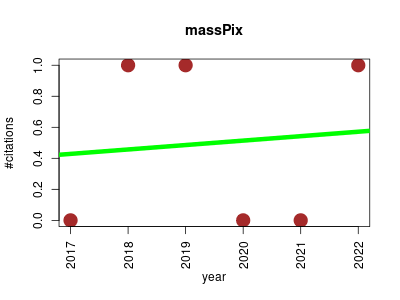 | 3 |
| ifindCPcli | Endocrinology and metabolism Enzymes Metabolomics Molecular interactions, pathways and networks Small molecules | findCPcli is a command line python-tool for the computation of chokepoint reactions in genome-scale metabolic models. The main purpose of the tool is to compute chokepoints by taking into account both the topology and the dynamic information of the network. In addition to the computation of chokepoints, findCPcli can compute and remove dead-end metabolites, find essential reactions and update the flux bounds of the reactions according to the results of Flux Variability Analysis. | 2021-10 | 3 | 1.5 | 3.0 | 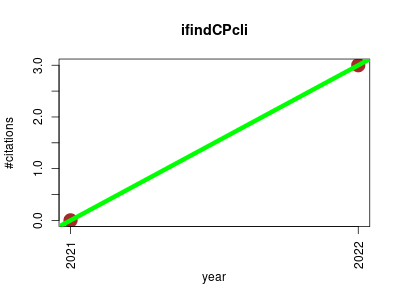 | 0 |
| iCYP-MFE | Drug discovery Drug metabolism Endocrinology and metabolism Metabolomics Small molecules | Identifying Human Cytochrome P450 Inhibitors Using Multitask Learning and Molecular Fingerprint-Embedded Encoding. | 2021-01 | 3 | 1.5 | 3.0 | 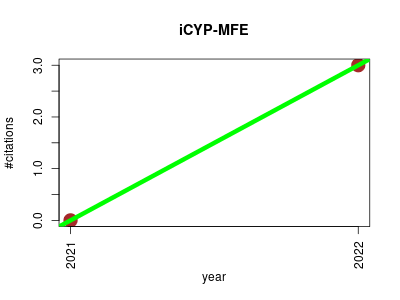 | 0 |
| MetEx | Metabolomics Proteomics Proteomics experiment Small molecules | Metabolomics Explorer Application for Natural Product Discovery. | 2021-12 | 3 | 1.5 | 3.0 | 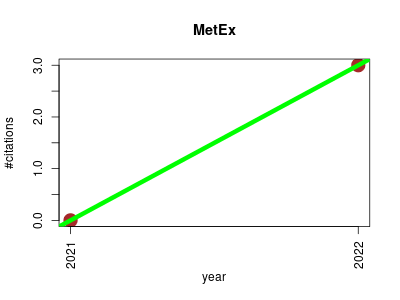 | 0 |
| TrpNet | Endocrinology and metabolism Metabolomics Microbial ecology Molecular interactions, pathways and networks Small molecules | TrpNet is a comprehensive resource for researchers to visually explore, search, or predict tryptophan metabolism within the context of human and mouse gut microbiome. | 2022-01 | 3 | 3.0 | 3.0 | 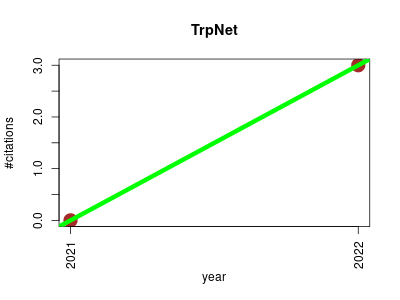 | 0 |
| NeatMS | Machine learning Metabolomics Proteomics experiment Workflows | NeatMS is an open source python package for untargeted LCMS signal labelling and filtering. NeatMS enables automated filtering of false positive MS peaks reported by commonly used LCMS data processing pipelines. NeatMS relies on neural network based classification. | 2022-03 | 3 | 3.0 | 0.0 | 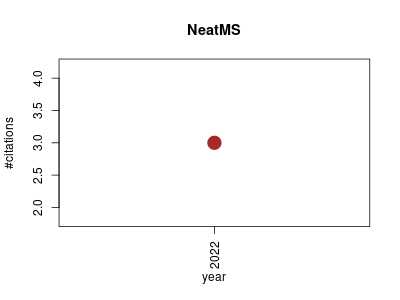 | 0 |
| MSNovelist | Lipids Machine learning Metabolomics Proteomics experiment Small molecules | De novo structure generation from mass spectra. | 2022-07 | 3 | 3.0 | 0.0 | 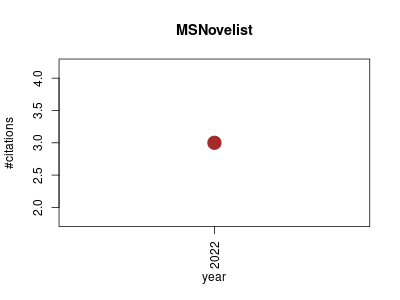 | 0 |
| LCMS TNO DECO | Metabolomics | A generic and automatic pre-processing tool for high resolution (HR) LC-MS and GC-MS data. | 2013-05 | 2 | 0.2 | -0.0 | 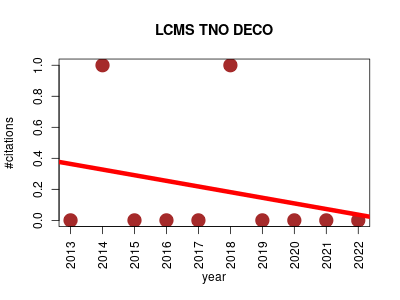 | 0 |
| BioSMXpress | Biochemistry Metabolomics | BiDesigned and developed as an enhancement to BioSM. It is, on average, 8 times faster than BioSM without compromising the quality of the predictions made. This software will be an extremely useful tool in the timely identification of unknown biochemical structures in metabolomics. | 2015-03 | 2 | 0.2 | 0.0 | 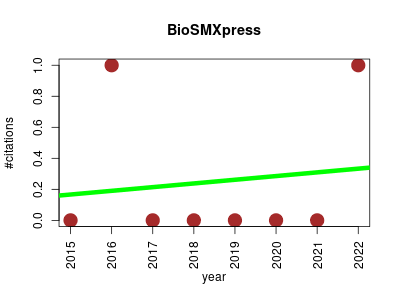 | 0 |
| MetMatch | Metabolomics | MetMatch is a software tool for the semi-automated comparison of different LC-HRMS chromatograms of untargeted metabolomics experiments. It provides methods to correct the raw-LC-HRMS data for m/z and retention time shifts relative to one or several preprocessed reference chromatograms. | 2016-12 | 2 | 0.3 | 0.1 | 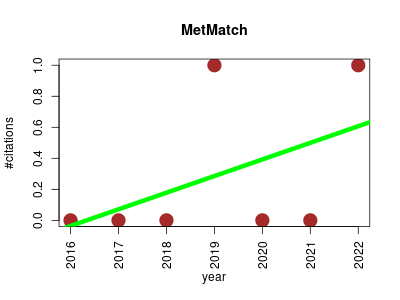 | 0 |
| msalign2 | Metabolomics Proteomics Proteomics experiment | Aligns 2 CE-MS or LC-MS datasets using accurate mass information. | 2009-12 | 2 | 0.1 | 0.0 | 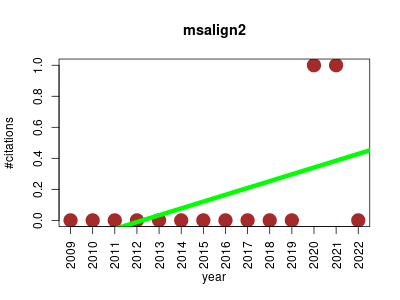 | 0 |
| MetTailor | Metabolomics Molecular interactions, pathways and networks | Software package that performs post-extraction processing steps such as cross-sample realignment and data normalization that are specifically designed to account for the experimental factors from chromatographic separation and MS analysis. | 2015-05 | 2 | 0.2 | -0.0 | 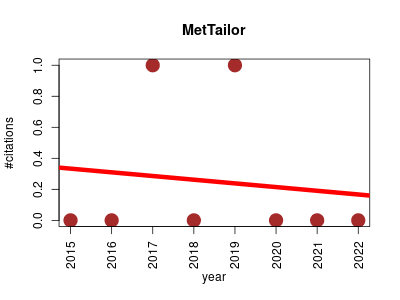 | 2 |
| mzExtract | Metabolomics | A new algorithm that enables untargeted metabolomics integration of samples that are measured with high resolution LC-MS. | 2013-11 | 2 | 0.2 | -0.0 | 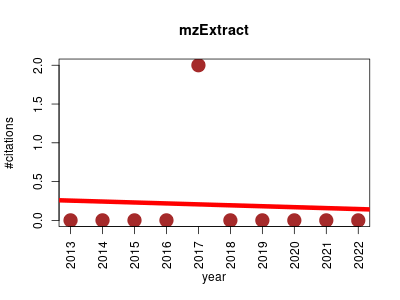 | 0 |
| multiview | Data mining Metabolomics | Software package for multiview pattern recognition methods. | 2019-08 | 2 | 0.5 | 0.0 | 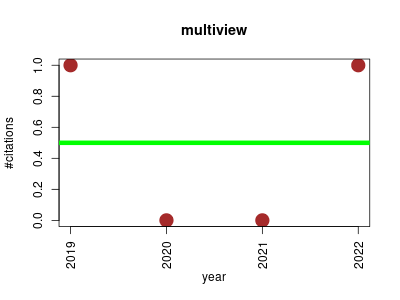 | 0 |
| Isoform | Fluxomics Metabolomics | R-program that prepares the corrected mass spectra (supposed to be provided by Midcor) to the final step of the workflow of fluxomic analysis: simulation of dynamics of mass isotopomer distribution in a kinetic model. | 2020-01 | 2 | 0.7 | 1.0 | 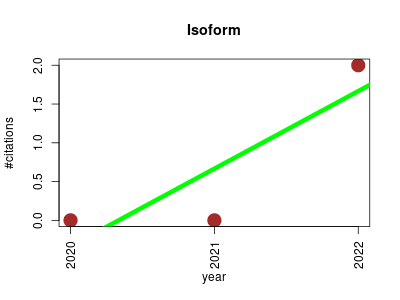 | 1 |
| Ramid | Fluxomics Metabolomics | R-program supporting the initial step of the workflow of 13C assisted fluxomic analysis: extraction of mass spectra (MS) of 13C-labeled metabolites of interest from raw mass spectrometer recordings saved in NetCDF files. | 2020-01 | 2 | 0.7 | 1.0 |  | 1 |
| LeGOO | Metabolomics Molecular interactions, pathways and networks Plant biology Proteomics Transcriptomics | An Expertized Knowledge Database for the Model Legume Medicago truncatula. | 2020-01 | 2 | 0.7 | 0.0 | 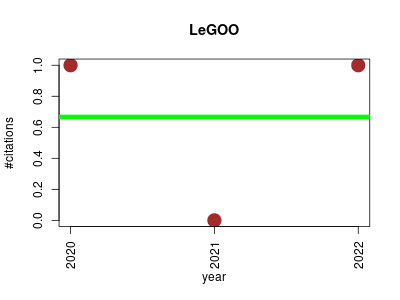 | 1 |
| MathIOmica | Metabolomics Personalised medicine Proteomics RNA-Seq Transcriptomics | General Analysis Utilities for Dynamic Omics Datasets. | 2020-03 | 2 | 0.7 | -0.0 | 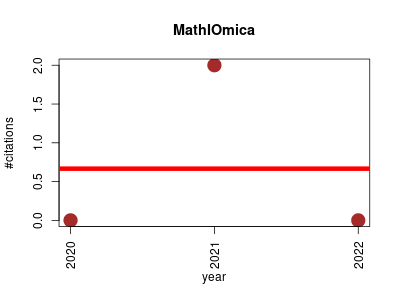 | 0 |
| way2drug | Endocrinology and metabolism Metabolomics Small molecules | MEDIUM CONFIDENCE! | CORRECT NAME OF TOOL COULD ALSO BE MetaTox | PASS-based prediction of metabolites detection in biological systems | Metabolite-Likeness No limit PaPi Pa0.5 Pa0.7 Pa0.9 | Predict metabolite for drawn structure | 2019-10 | 2 | 0.5 | 0.2 | 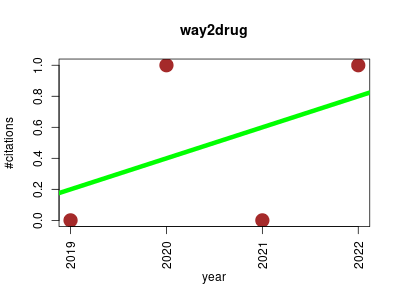 | 0 |
| PATHsupcre8 | Endocrinology and metabolism Enzymes Metabolomics Molecular interactions, pathways and networks Small molecules | A Tool That Facilitates the Searching for Heterologous Biosynthetic Routes. | 2020-12 | 2 | 0.7 | 0.5 | 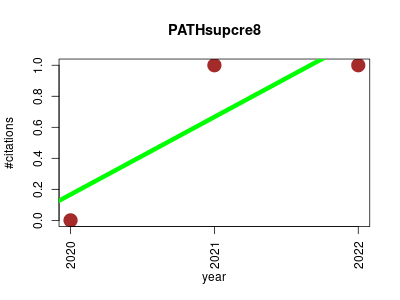 | 1 |
| liputils | Endocrinology and metabolism Informatics Lipids Metabolomics Proteomics experiment | A Python module to manage individual fatty acid moieties from complex lipids. | 2020-12 | 2 | 0.7 | 0.5 | 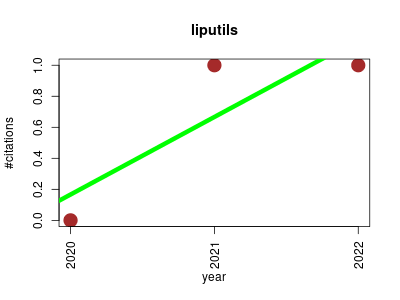 | 1 |
| SPECS | Biomarkers Metabolomics Protein structure analysis RNA-Seq Structure prediction | a non-parametric method to identify tissue-specific molecular features for unbalanced sample groups. | 2020-02 | 2 | 0.7 | -1.0 | 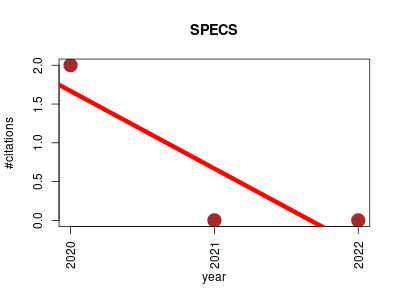 | 0 |
| MRMkit | Analytical chemistry Metabolomics Proteomics Proteomics experiment Workflows | Automated Data Processing for Large-Scale Targeted Metabolomics Analysis. | 2020-10 | 2 | 0.7 | 0.5 | 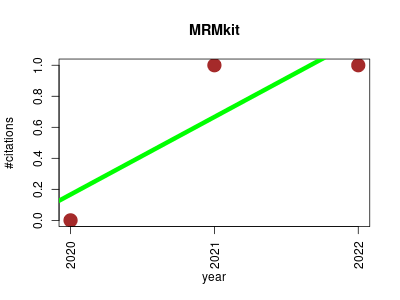 | 1 |
| ncGTW | Metabolomics Proteomics Proteomics experiment Sequence analysis | Targeted realignment of LC-MS profiles by neighbor-wise compound-specific graphical time warping with misalignment detection. | 2020-05 | 2 | 0.7 | 0.5 | 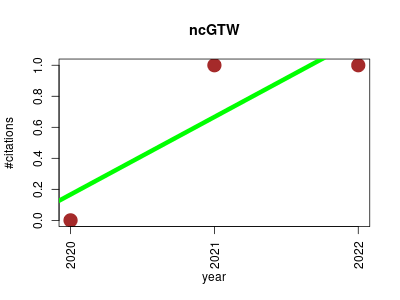 | 1 |
| DaDIA | Metabolomics | DaDIA is a LC-MS feature extraction and annotation workflow in R. | 2021-02 | 2 | 1.0 | 2.0 | 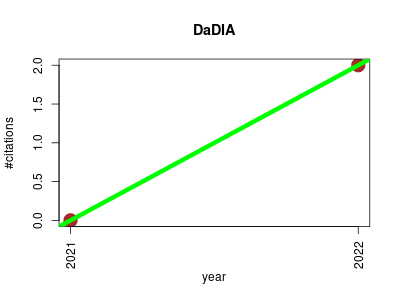 | 0 |
| LM-GlycomeAtlas | Biochemistry Drug discovery Gene expression Imaging Metabolomics Microarray experiment Proteomics | A Novel Visualization Tool for Lectin Microarray-Based Glycomic Profiles of Mouse Tissue Sections | LM-GlycomeAtlas v.1.0 is a web tool visualizing the data from Lectin Microarray analyses using 45 lectins by the Kuno Laboratory at AIST | This work was supported by a project for utilizing glycans in the development of innovative drug discovery technologies in the project focused on developing key technology for discovering and manufacturing drugs for next-generation treatment and diagnosis from the Japan Agency for Medical Research and Development (AMED) | CFG(Consortium for Functional Glycomics) Glycan Profilling data | Mapping data of human serum data was from Dr. Shunji Natsuka of Niigata University, Japan | 2019-08 | 2 | 0.5 | 0.2 | 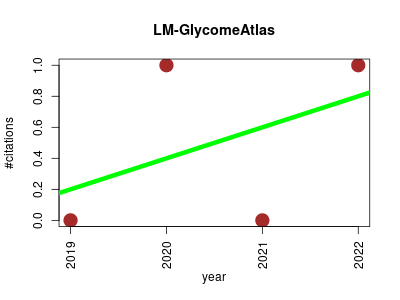 | 0 |
| leapR | Endocrinology and metabolism Metabolomics Molecular interactions, pathways and networks Proteomics Sequence analysis | An R Package for Multiomic Pathway Analysis | 2021-04 | 2 | 1.0 | 2.0 | 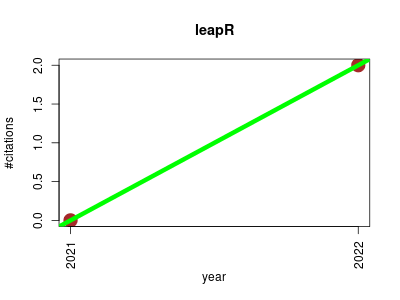 | 1 |
| COMETS Analytics | Bioinformatics Metabolomics Public health and epidemiology Small molecules Statistics and probability | COMETS Analytics is an online tool for analyzing and meta-analyzing metabolomics data in large research consortia. | -- | 2 | 0.0 | 0.0 | 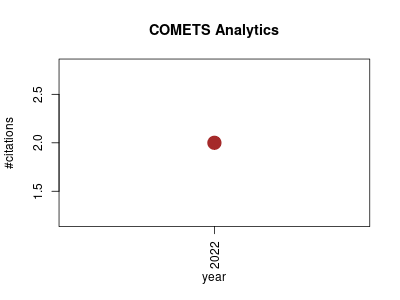 | 1 |
| CyProduct | Cheminformatics Endocrinology and metabolism Enzymes Metabolomics Small molecules | CyProduct is a software tool for accurately predicting the byproducts of human Cytochrome P450 metabolism. | 2021-06 | 2 | 1.0 | 2.0 | 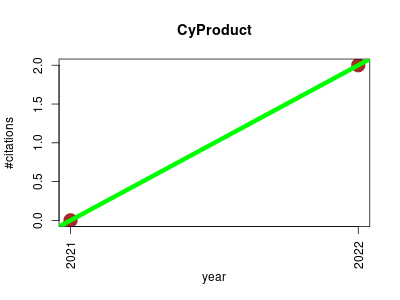 | 0 |
| GeenaR | Metabolomics Oncology Proteomics Proteomics experiment Sequence analysis | GeenaR is a tool for MALDI/ToF MS spectra analysis. | -- | 2 | 0.0 | 2.0 |  | 1 |
| metabCombiner | Metabolomics Proteomics Proteomics experiment Small molecules | metabCombiner is a R package for paired untargeted LC-HR-MS metabolomics feature matching and concatenation of disparately acquired datasets. | 2021-03 | 2 | 1.0 | 0.0 |  | 0 |
| mwTab | Metabolomics NMR Proteomics Proteomics experiment Small molecules | mwTab is a Python library for RESTful access and enhanced quality control, deposition, and curation of the metabolomics workbench data repository. The mwtab package is a Python library that facilitates reading and writing files in mwTab format used by the Metabolomics Workbench for archival of Mass Spectrometry (MS) and Nuclear Magnetic Resonance (NMR) experimental data. | 2021-03 | 2 | 1.0 | 2.0 |  | 1 |
| NetGSA | Endocrinology and metabolism Gene expression Metabolomics Molecular interactions, pathways and networks | Carry out network-based gene set analysis by incorporating external information about interactions among genes, as well as novel interactions learned from data. | 2021-06 | 2 | 1.0 | 2.0 |  | 0 |
| MassBase | Endocrinology and metabolism Metabolomics Plant biology Proteomics experiment Small molecules | A large-scaled depository of mass spectrometry datasets for metabolome analysis. | 2021-01 | 2 | 1.0 | 0.0 |  | 1 |
| POMAShiny | Biomarkers Machine learning Metabolomics Proteomics Sequence analysis Workflows | POMAShiny is an user-friendly web-based workflow for metabolomics and proteomics data analysis. | 2021-07 | 2 | 1.0 | 2.0 |  | 0 |
| SMILE | Endocrinology and metabolism Evolutionary biology Machine learning Metabolomics Small molecules | Systems Metabolomics using Interpretable Learning and Evolution (SMILE) is a framework for for supervised metabolomics data analysis. SMILE package implements Linear Genetic Programming (LGP) algorithm in python, with a scikit-learn style API. It is mainly used in data mining and finding feature interactions. Note it currently only support binary classification data. | 2021-12 | 2 | 1.0 | 2.0 |  | 0 |
| ADAP-KDB | Analytical chemistry Metabolomics Proteomics experiment Workflows | A Spectral Knowledgebase for Tracking and Prioritizing Unknown GC-MS Spectra in the NIHs Metabolomics Data Repository. | 2021-09 | 2 | 1.0 | 2.0 |  | 1 |
| R2DGC | Analytical chemistry Endocrinology and metabolism Metabolomics Proteomics experiment | Provides functions for aligning 2D gas chromatography mass spectrometry derived metabolite peaks obtained from primary processing and generates an alignment table that allows for a comparison of common peaks across samples and metabolite identification. | 2018-05 | 2 | 0.4 | 0.1 |  | 1 |
| mapbayr | Drug metabolism Metabolomics Small molecules Statistics and probability | Easy and reliable maximum a posteriori Bayesian estimation of pharmacokinetic parameters with the open-source R package mapbayr. | 2021-10 | 2 | 1.0 | 2.0 |  | 0 |
| BiG-MAP | Endocrinology and metabolism Metabolomics Metagenomics Metatranscriptomics Microbial ecology | The Biosynthetic Gene cluster Meta’omics abundance Profiler (BiG-MAP). A command-line tool that it is able to profile the abundance and expression of a collection of gene clusters across metagenomic and metatranscriptomic data from any kind of biome, including human, plant, animal, marine, and soil microbiomes. | 2021-09 | 2 | 1.0 | 2.0 |  | 0 |
| MIMOSA2 | Endocrinology and metabolism Metabolomics Microbial ecology Molecular interactions, pathways and networks Small molecules | MIMOSA2 is a tool for metabolic model-based evaluation of paired microbiome and metabolomics datasets. MIMOSA2 1) constructs community metabolic models, 2) assesses whether metabolite measurements are consistent with estimated community metabolic potential, and 3) identifies specific taxa and reactions that can explain metabolite variation. | -- | 2 | 0.0 | 0.0 |  | 1 |
| FunOrder | Mapping Metabolomics Model organisms Phylogenetics Proteomics | A robust and semi-automated method for the identification of essential biosynthetic genes through computational molecular co-evolution. | 2021-09 | 2 | 1.0 | 2.0 |  | 1 |
| CRISP | Imaging Machine learning Metabolomics Proteomics experiment Urology and nephrology | CRISP: a deep learning architecture for GC × GC–TOFMS contour ROI identification, simulation and analysis in imaging metabolomics | 2022-03 | 2 | 2.0 | 0.0 |  | 0 |
| NP Analyst | Compound libraries and screening Metabolomics Proteomics experiment Small molecules Workflows | NP Analyst is a platform-independent tool for integrating bio-activity data with metabolomics data. | 2022-02 | 2 | 2.0 | 0.0 |  | 0 |
| OncoboxPD | Endocrinology and metabolism Gene expression Metabolomics Molecular interactions, pathways and networks Small molecules | Human 51 672 molecular pathways database with tools for activity calculating and visualization. | 2022-01 | 2 | 2.0 | 0.0 |  | 0 |
| RareLSD | DNA polymorphism Enzymes Metabolomics Rare diseases Small molecules | a manually curated database of lysosomal enzymes associated with rare diseases. | 2019-01 | 2 | 0.5 | 0.0 |  | 0 |
| rMSIKeyIon | Metabolomics Proteomics experiment Statistics and probability | Ion Filtering R Package for Untargeted Analysis of Metabolomic LDI-MS Images. | 2019-08 | 1 | 0.2 | -0.1 |  | 1 |
| MetPC | Biomarkers Metabolomics Small molecules | Metabolite Pipeline Consisting of Metabolite Identification and Biomarker Discovery Under the Control of Two-Dimensional FDR. | 2019-05 | 1 | 0.2 | 0.1 | 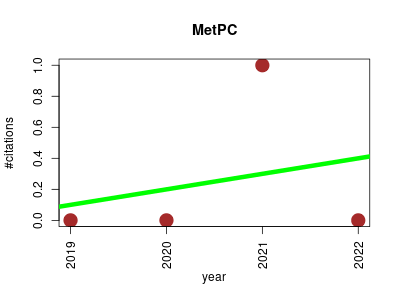 | 0 |
| SBM | Gene transcripts Metabolomics Molecular interactions, pathways and networks | Analysis of correlation-based biomolecular networks from different omics data by fitting stochastic block models | SBM-for-correlation-based-networks | 2019-01 | 1 | 0.2 | -0.1 | 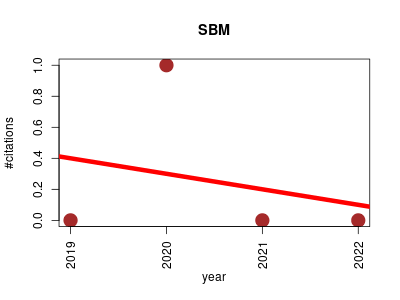 | 0 |
| nonadherence | Haematology Metabolomics Small molecules | Prediction of Thiopurine Metabolite Levels Based on Haematological and Biochemical Parameters. | 2019-10 | 1 | 0.2 | 0.1 | 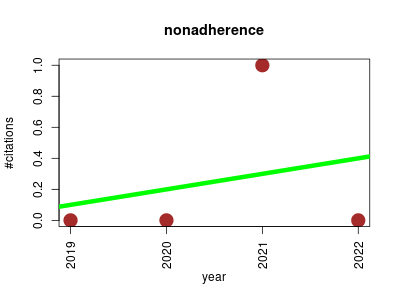 | 0 |
| BRAT | Biomarkers Drug metabolism Metabolomics Public health and epidemiology Small molecules | The Biomarker Reliability Assessment Tool (BRAT) was developed to help researchers set targets to improve exposure assessment in studies of chemicals with a short half-life. The BRAT software uses a simple pharmacokinetic model to simulate exposure, internal levels, and resulting urinary levels in individuals from a population based on user-specified inputs (e.g., biological half-life, within- and between-person variability in exposure). Based on internal levels (a toxicologically-relevant exposure metric) and urinary levels in the simulated population, the tool evaluates – through linear regression and quantile classification – the accuracy of the estimation of internal levels depending on the number of urine samples. | 2020-12 | 1 | 0.3 | 0.5 | 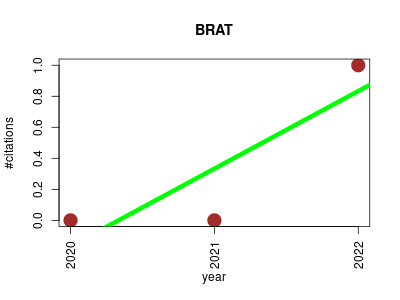 | 1 |
| JUMPm | Chemistry Metabolomics Model organisms Proteomics experiment Small molecules | A Tool for Large-Scale Identification of Metabolites in Untargeted Metabolomics. | 2020-05 | 1 | 0.3 | 0.5 | 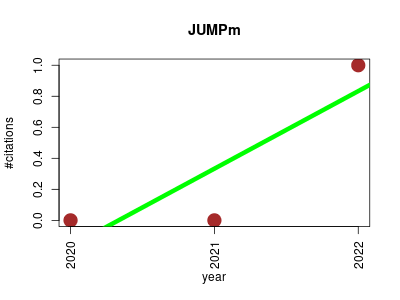 | 1 |
| MetaPASS | Drug discovery Endocrinology and metabolism Metabolomics Pharmacology Small molecules | A Web Application for Analyzing the Biological Activity Spectrum of Organic Compounds Taking into Account their Biotransformation. | 2021-04 | 1 | 0.5 | -1.0 | 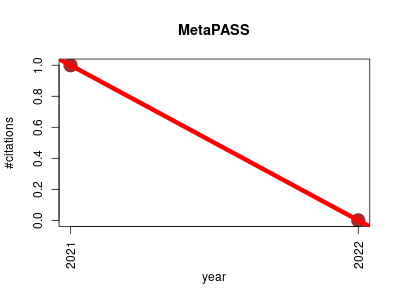 | 1 |
| SUMMER | Endocrinology and metabolism Enzymes Metabolomics Molecular interactions, pathways and networks Small molecules | shiny utility for metabolomics and multiomics exploratory research. | 2020-05 | 1 | 0.3 | -0.0 | 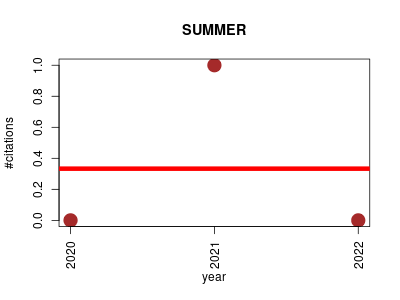 | 1 |
| SWIFTCORE | Endocrinology and metabolism Fluxomics Metabolomics Molecular interactions, pathways and networks Small molecules | a tool for the context-specific reconstruction of genome-scale metabolic networks. | 2020-04 | 1 | 0.3 | -0.0 | 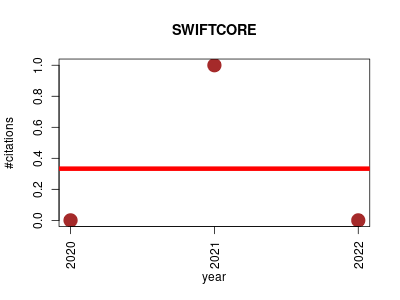 | 1 |
| MMHub | Biological databases Metabolomics Plant biology Proteomics experiment Small molecules | Database for the mulberry metabolome. | 2020-01 | 1 | 0.3 | 0.5 | 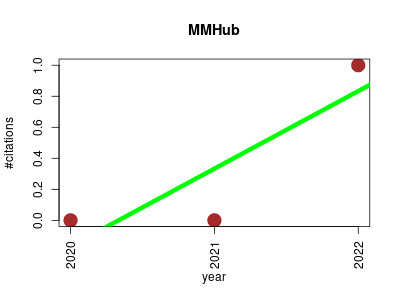 | 0 |
| MSpectraAI | Machine learning Metabolomics Proteomics Proteomics experiment Sequence analysis | A powerful platform for deciphering proteome profiling of multi-tumor mass spectrometry data by using deep neural networks. | 2020-10 | 1 | 0.3 | 0.5 | 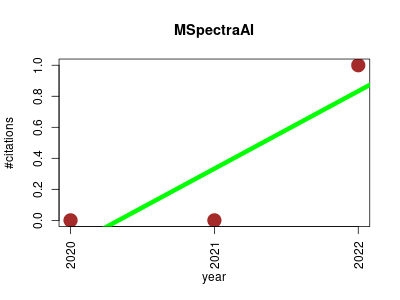 | 1 |
| ToxinDB | Cheminformatics Enzymes Metabolomics Small molecules Systems biology | A data-driven integrative platform for computational prediction of toxin biotransformation with a case study. | 2021-04 | 1 | 0.5 | 1.0 | 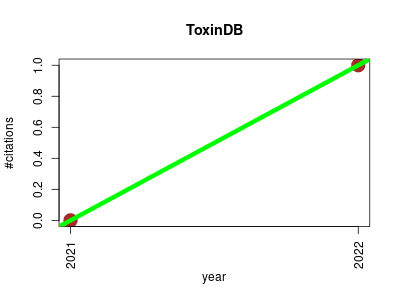 | 0 |
| EpiMetal | Endocrinology and metabolism Lipids Mapping Metabolomics Public health and epidemiology | EpiMetal is an open-source graphical web browser tool for easy statistical analyses in epidemiology and metabolomics. | 2020-08 | 1 | 0.3 | -0.0 | 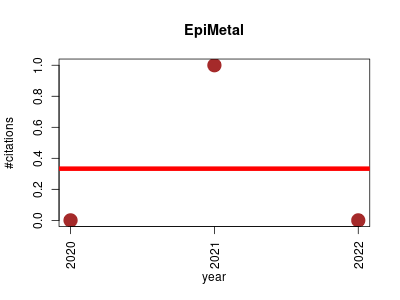 | 1 |
| OmicsON | Lipids Metabolomics Proteomics Statistics and probability Transcriptomics | Integration of omics data with molecular networks and statistical procedures. | 2020-07 | 1 | 0.3 | 0.5 | 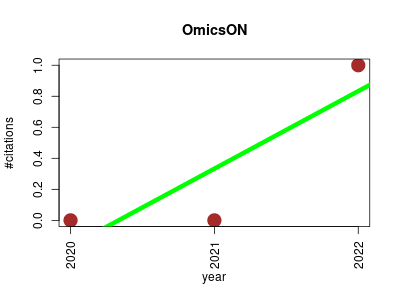 | 1 |
| CANTARE | Metabolomics Metagenomics Microbial ecology Molecular interactions, pathways and networks Workflows | CANTARE (Consolidated Analysis of Network Topology And Regression Elements) is a workflow for building predictive regression models from network neighborhoods in multi-omic networks. | 2021-12 | 1 | 0.5 | 1.0 | 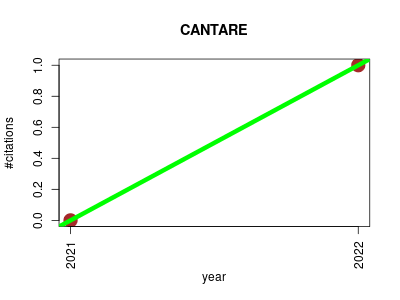 | 0 |
| OpenTIMS | Lipids Metabolomics Proteomics Proteomics experiment Sequence analysis | Open and Easy Access to timsTOF Raw Data. | 2021-04 | 1 | 0.5 | 1.0 |  | 0 |
| lcmsWorld | Imaging Metabolomics Proteomics Proteomics experiment Sequence analysis | High-Performance 3D Visualization Software for Mass Spectrometry. | 2021-04 | 1 | 0.5 | 1.0 | 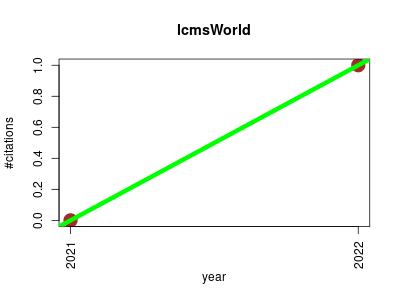 | 0 |
| gcProfileMakeR | Genotype and phenotype Machine learning Metabolomics Physiology Small molecules | gcProfileMakeR is an R package for automatic classification of constitutive and non-constitutive metabolites. | -- | 1 | 0.0 | -1.0 | 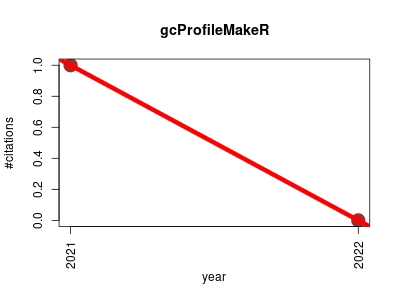 | 0 |
| Xconnector | Biological databases Metabolomics Molecular interactions, pathways and networks Small molecules | Xconnector is a software package designed to easily retrieve, and visualize metabolomics data from different database sources. | 2021-08 | 1 | 0.5 | 1.0 |  | 0 |
| mzRAPP | Metabolomics Proteomics experiment Workflows | mzRAPP is a tool for reliability assessment of data pre-processing in non-targeted metabolomics. The goal of mzRAPP is to allow reliability assessment of non-targeted data pre-processing (NPP; XCMS, XCMS3, MetaboanalystR 3.0, XCMS-online, MZmine 2, MS-DIAL, OpenMS, El-MAVEN,..) in the realm of liquid chromatography high-resolution mass spectrometry (LC-HRMS). mzRAPPs approach is based on the increasing popularity of merging non-targeted with targeted metabolomics meaning that both types of data evaluation are often performed on the same dataset. | -- | 1 | 0.0 | 1.0 |  | 0 |
| mWISE | Metabolomics Proteomics experiment Small molecules | An Algorithm for Context-Based Annotation of Liquid Chromatography-Mass Spectrometry Features through Diffusion in Graphs. | 2021-08 | 1 | 0.5 | 1.0 |  | 0 |
| NICEdrug.ch | Drug metabolism Endocrinology and metabolism Medicinal chemistry Metabolomics Small molecules | NICEdrug.ch is a resource allowing systematic and large-scale computational analysis of drug biochemistry (metabolic precursors or prodrugs and metabolic fate or degradation), enzymatic targets, and toxicity in the context of cellular metabolism, i.e. currently including: human, Plasmodium (malaria parasite), and E. coli metabolism. | 2021-01 | 1 | 0.5 | 1.0 |  | 0 |
| SimExTargId | Data acquisition Metabolomics Proteomics experiment Structural biology | Comprehensive package for real-time LC-MS data acquisition and analysis. | 2018-10 | 1 | 0.2 | -0.1 |  | 1 |
| IMFLer | Endocrinology and metabolism Fluxomics Metabolomics Molecular interactions, pathways and networks Small molecules | A Web Application for Interactive Metabolic Flux Analysis and Visualization. | 2021-10 | 1 | 0.5 | 1.0 |  | 0 |
| QSDB | Metabolomics Microbial ecology Model organisms Small molecules Systems biology | Quorumpeps® is a resource of quorum sensing signalling peptides. This database is managed by Ghent University. Based upon your input, this search page will give you all information (structure, activity, physicochemical properties and related literature). The database is linked to a manuscript entitled Quorumpeps database: chemical space, microbial origin and functionality of quorum sensing peptides, in which the origin of the different peptides and their quorum sensing pathways and methods are described. | 2021-01 | 1 | 0.5 | 1.0 |  | 0 |
| MaRR | Metabolomics Microarray experiment Proteomics experiment Public health and epidemiology Small molecules | This app provides a user interface for the open-source R package marr, which implements the method Maximum Rank Reproducibility (MaRR), a nonparametric approach that detects reproducible signals using a maximal rank statistic for high-dimensional biological data. | 2021-12 | 1 | 0.5 | 1.0 |  | 0 |
| SLAW | Computer science Lipids Metabolomics Proteomics experiment Workflows | A Scalable and Self-Optimizing Processing Workflow for Untargeted LC-MS. | 2021-11 | 1 | 0.5 | 1.0 |  | 0 |
| PROMISed | Metabolomics Plant biology Proteomics Proteomics experiment Small molecules | A novel web-based tool to facilitate analysis and visualization of the molecular interaction networks from co-fractionation mass spectrometry (CF-MS) experiments. | 2021-01 | 1 | 0.5 | 1.0 |  | 0 |
| dbPUP | Endocrinology and metabolism Gene and protein families Metabolomics Metagenomics Microbial ecology | dbPUP is a database of polyphenol utilization proteins (PUPs) that have been experimentally characterized to metabolize polyphenol substrates. | 2022-02 | 1 | 1.0 | 0.0 |  | 0 |
| LCMD | Biomarkers Metabolomics Oncology Proteomics experiment Small molecules | The Lung Cancer Metabolome Database (LCMD), a freely available online database depositing 2013 lung cancer-related metabolites identified from 65 mass spectrometry-based lung cancer metabolomics studies. | 2022-01 | 1 | 1.0 | 0.0 |  | 1 |
| DEIMoS | Metabolomics NMR Proteomics experiment Workflows | DEIMoS: Data Extraction for Integrated Multidimensional Spectrometry, a Python application programming interface (API) and command-line tool for high-dimensional mass spectrometry data analysis workflows that offers ease of development and access to efficient algorithmic implementations. | 2022-04 | 1 | 1.0 | 0.0 |  | 0 |
| PaintOmics | Functional, regulatory and non-coding RNA Metabolomics Molecular interactions, pathways and networks Plant biology Small molecules | New tools for the integrative analysis of multi-omics datasets supported by multiple pathway databases. | 2022-07 | 1 | 1.0 | 0.0 |  | 0 |
| hECA | Cell biology Human biology Metabolomics Regenerative medicine Sequence assembly | The human Ensemble Cell Atlas (hECA) a database. | 2022-05 | 1 | 1.0 | 0.0 |  | 0 |
| IDSL-IPA | Metabolomics Model organisms Proteomics Proteomics experiment | IDSL.IPA Characterizes the Organic Chemical Space in Untargeted LC/HRMS Data Sets. | 2022-06 | 1 | 1.0 | 0.0 |  | 0 |
| PSAMM | Metabolomics Molecular interactions, pathways and networks | Open source software that is designed for the curation and analysis of metabolic models. It supports model version tracking, model annotation, data integration, data parsing and formatting, consistency checking, automatic gap filling, and model simulations. | 2018-02 | 0 | 0.0 | 0.0 |  | 0 |
| MetaboCraft | Data visualisation Metabolomics Molecular interactions, pathways and networks | Minecraft plugin which creates immersive visualizations of metabolic networks and pathways in a 3D environment and allows the results of user experiments to be viewed in this context, presenting a novel approach to exploring the metabolome. | 2018-08 | 0 | 0.0 | 0.0 |  | 0 |
| G.A.M.E | Cheminformatics Computational chemistry Metabolomics Software engineering | GPU-Accelerated Mixture Elucidator. GPU acceleration is useful in solving complex chemical information problems. Identifying unknown structures from the mass spectra of natural product mixtures has been a desirable yet unresolved issue in metabolomics. | 2017-09 | 0 | 0.0 | 0.0 |  | 0 |
| mSpecs Editor | Metabolomics Proteomics experiment | Free tool for managing mass spectral libraries with an additional focus on gas chromatography / mass spectrometry, liquid chromatography / mass spectrometry and related techniques. | 2009-07 | 0 | 0.0 | 0.0 |  | 0 |
| RDFIO | Data management Metabolomics Rare diseases | Extends the RDF import and export functionality in Semantic MediaWiki (SMW) to enable using SMW as a collaborative platform for editing semantic data. | 2017-09 | 0 | 0.0 | 0.0 |  | 0 |
| TRC | Metabolomics Proteomics experiment | Truncated rank correlation (TRC) as a robust measure of test-retest reliability in mass spectrometry data. | 2019-01 | 0 | 0.0 | 0.0 |  | 0 |
| BCSExplorer | Compound libraries and screening Enzymes Metabolomics Molecular biology Small molecules | A Customized Biosynthetic Chemical Space Explorer with Multifunctional Objective Function Analysis. | 2020-03 | 0 | 0.0 | 0.0 |  | 0 |
| HastaLaVista | Biomedical science Endocrinology and metabolism Metabolomics Small molecules Surgery | Web-based user interface for NMR-based untargeted metabolic profiling analysis in biomedical sciences. | 2019-12 | 0 | 0.0 | 0.0 |  | 0 |
| CAMND | Endocrinology and metabolism Metabolomics Small molecules | Comparative analysis of metabolic network decomposition based on previous and two new criteria. | 2020-03 | 0 | 0.0 | 0.0 |  | 0 |
| CardiacPBPK | Drug metabolism Metabolomics Physiology Safety sciences Small molecules | A tool for the prediction and visualization of time-concentration profiles of drugs in heart tissue. | 2019-12 | 0 | 0.0 | 0.0 |  | 0 |
| LLCT | Genotype and phenotype Mapping Metabolomics Microarray experiment Proteomics | Longitudinal linear combination test for gene set analysis. | 2019-12 | 0 | 0.0 | 0.0 |  | 0 |
| MetaMed | Medicine Metabolomics Metagenomics Microbial ecology Small molecules | Linking Microbiota Functions with Medicine Therapeutics. | 2019-01 | 0 | 0.0 | 0.0 |  | 0 |
| FlavoDb | Drug discovery Literature and language Metabolomics Nutritional science Small molecules | A web-based chemical repository of flavonoid compounds. | 2019-11 | 0 | 0.0 | 0.0 |  | 0 |
| Atlas of Inflammation-Resolution (AIR) | Allergy, clinical immunology and immunotherapeutics Genotype and phenotype Metabolomics Molecular interactions, pathways and networks Pathology | The Atlas of Inflammation-Resolution (AIR) is a web application for capturing an essential part ofacute inflammation and inflammation resolution research. | -- | 0 | 0.0 | 0.0 |  | 0 |
| ichespa | Compound libraries and screening Metabolomics Molecular biology Proteomics experiment Small molecules | Streamlining Expansive Chemical Space Evaluation of Molecular Sets. | 2020-12 | 0 | 0.0 | 0.0 |  | 0 |
| MixONat | Metabolomics NMR Nutritional science Proteomics experiment Small molecules | Software for the Dereplication of Mixtures Based on 13C NMR Spectroscopy. | 2020-07 | 0 | 0.0 | 0.0 |  | 0 |
| ExpoKids | Ecology Imaging Literature and language Metabolomics Public health and epidemiology | ExpoKids is an R-based tool for characterizing aggregate chemical exposure during childhood. | 2021-03 | 0 | 0.0 | 0.0 |  | 0 |
| WHONDRS | Metabolomics Metagenomics Metatranscriptomics Microbial ecology Small molecules | a web application for global survey of surface water metabolites. | 2020-07 | 0 | 0.0 | 0.0 |  | 0 |
| M2R | Endocrinology and metabolism Genotype and phenotype Metabolomics Physiology Small molecules | A Python add-on to cobrapy for modifying human genome-scale metabolic reconstruction using the gut microbiota models. | -- | 0 | 0.0 | 0.0 |  | 0 |
| MetaRon | Gene structure Metabolomics Metagenomics Microbial ecology Small molecules | A pipeline for prediction of Metagenome and whole-genome opeRons. | 2021-12 | 0 | 0.0 | 0.0 |  | 0 |
| Fantastic databases and where find them | Biological databases Epigenetics Functional, regulatory and non-coding RNA Metabolomics Proteomics | Fantastic databases and where find them is a collection of databases for omics data. | -- | 0 | 0.0 | 0.0 |  | 0 |
| ASTHERISC | Biotechnology Endocrinology and metabolism Enzymes Metabolomics Small molecules | ASTHERISC (Algorithmic Search of THERmodynamic advantages In stoichiometric Single-species Community models). Designing microbial communities to maximize the thermodynamic driving force for the production of chemicals. | 2021-06 | 0 | 0.0 | 0.0 |  | 0 |
| RmsiGUI | Imaging Metabolomics Proteomics experiment Small molecules | RmsiGUI provides a graphical user interface (GUI) to analyze mass spectrometry imaging (MSI) data with the statistical language R. The interface is based on shiny and visualized in a web browser. | 2021-01 | 0 | 0.0 | 0.0 |  | 0 |
| TCM-Blast | Medicine Metabolomics Molecular interactions, pathways and networks Plant biology Whole genome sequencing | TCM-Blast for traditional Chinese medicine genome alignment with integrated resources. | 2021-12 | 0 | 0.0 | 0.0 |  | 0 |
| SCOUR | Fluxomics Machine learning Metabolomics Molecular interactions, pathways and networks Small molecules | A stepwise machine learning framework for predicting metabolite-dependent regulatory interactions. | 2021-12 | 0 | 0.0 | 0.0 |  | 0 |
| SGI | Biochemistry Metabolomics Proteomics | The Subgroup Identification (SGI) toolbox provides an algorithm to automatically detect clinical subgroups of samples in large-scale omics datasets. It is based on hierarchical clustering trees in combination with a specifically designed association testing and visualization framework that can process a large number of clinical parameters and outcomes in a systematic fashion. A multi-block extension allows for the simultaneous use of multiple omics datasets on the same samples. | -- | 0 | 0.0 | 0.0 |  | 0 |
| MS2Planner | Data acquisition Metabolomics Molecular biology Proteomics experiment Small molecules | Optimizing Coverage in Tandem Mass Spectrometry based Metabolomics by Iterative Data Acquisition | 2021-07 | 0 | 0.0 | 0.0 |  | 0 |
| Amanida | Metabolomics Preclinical and clinical studies Small molecules | An R package for Meta-analysis of metabolomics non-integral data. | -- | 0 | 0.0 | 0.0 |  | 0 |
| MStractor | Metabolomics Model organisms Proteomics experiment Small molecules Workflows | MStractor is an R workflow package for non-targeted processing of LC-MS data. | 2021-08 | 0 | 0.0 | 0.0 |  | 0 |
| multiMarker | Biomarkers Metabolomics Nutritional science | multiMarker is a web application that infers the relationship between biomarker and food quantity data from an intervention study, and allows prediction of food intake when only biomarker data are available. Additionally, multiMarker provides quantification of the uncertainty in intake predictions. | 2021-12 | 0 | 0.0 | 0.0 |  | 0 |
| mspack | Metabolomics Proteomics Proteomics experiment | mspack is a C++ program for lossless and lossy mass spectrometry data compression, achieving a high compression ratio without sacrificing performance. It includes an example implementation for mzXML and mzML as well as a format-agnostic API. | -- | 0 | 0.0 | 0.0 |  | 0 |
| FiCoS | Enzymes Mathematics Metabolomics Molecular dynamics Molecular interactions, pathways and networks | FiCoS (Fine- and Coarse-grained Simulator) is a novel efficient deterministic simulator of biological systems based on two different integration methods belonging to the Runge-Kutta family: the Dormand–Prince (DOPRI) method [1, 2], used in the absence of stiffness, and the Radau IIA method [3, 4] exploited when the system is stiff. | 2021-09 | 0 | 0.0 | 0.0 |  | 0 |
| rMisbeta | Data acquisition Metabolomics Transcriptomics | A robust missing value imputation approach in transcriptomics and metabolomics data. | 2021-11 | 0 | 0.0 | 0.0 |  | 0 |
| MoNET | Biomarkers DNA polymorphism Metabolomics Molecular interactions, pathways and networks Small molecules | MoNET is an R package providing network analysis of -omics findings. It is built on top of an integrated network, including metabolite-protein interactions, protein interactions and relationships between genetic variations and transcription factor binding sites. | -- | 0 | 0.0 | 0.0 |  | 0 |
| fobitoolsGUI | Metabolomics Molecular interactions, pathways and networks Nutritional science Ontology and terminology Small molecules | This package provides a set of tools for interacting with FOBI (Food-Biomarker Ontology). | -- | 0 | 0.0 | 0.0 |  | 0 |
| XDeathDB | Cell biology Immunogenetics Metabolomics Molecular interactions, pathways and networks Oncology | A visualization platform for cell death molecular interactions. | 2021-12 | 0 | 0.0 | 0.0 |  | 0 |
| SpinSPJ | MRI Machine learning Metabolomics NMR Software engineering | A novel NMR scripting system to implement artificial intelligence and advanced applications. | 2021-12 | 0 | 0.0 | 0.0 |  | 0 |
| Aristotle | Genomics Metabolomics Personalised medicine Proteomics Sequence analysis | Aristotle addresses the two challenges intrinsic to omics data: high dimensionality and hidden stratification. It employs existing biological knowledge and a state-of-the-art patient stratification method to tackle the above challenges and applies a quasi-experimental design method to each stratum to find stratum-specific potential causes. | 2022-12 | 0 | 0.0 | 0.0 |  | 0 |
| BayesSUR | Biomarkers Genotype and phenotype Metabolomics NMR Small molecules | A computationally efficient Bayesian seemingly unrelated regressions model for high-dimensional quantitative trait loci discovery. | 2021-08 | 0 | 0.0 | 0.0 |  | 0 |
| RawHummus | Lipids Metabolomics Proteomics Proteomics experiment | An R Shiny app for assessing LC–MS system performance by visualising instrument log files and monitoring raw quality control samples within a project. | 2022-04 | 0 | 0.0 | 0.0 |  | 0 |
| PPS | Gynaecology and obstetrics Metabolomics Molecular interactions, pathways and networks Small molecules | R package to perform PPS analysis on graphical models | 2022-12 | 0 | 0.0 | 0.0 |  | 0 |
| MEMO | Metabolomics | Ms2 basEd saMple vectOrization (MEMO) is a method allowing a Retention Time (RT) agnostic alignment of metabolomics samples using the fragmentation spectra (MS2) of their consituents. The occurence of MS2 peaks and neutral losses (to the precursor) in each sample is counted and used to generate an MS2 fingerprint of the sample. These fingerprints can in a second stage be aligned to compare different samples. | -- | 0 | 0.0 | 0.0 |  | 0 |
| GeneCup | GWAS study Machine learning Metabolomics Natural language processing Ontology and terminology | GeneCup automatically extracts information from PubMed and NHGRI-EBI GWAS catalog on the relationship of any gene with a custom list of keywords hierarchically organized into an ontology. | 2022-05 | 0 | 0.0 | 0.0 |  | 0 |
| JPA | Endocrinology and metabolism Metabolomics Proteomics experiment Small molecules | JPA (short for Joint Metabolomics Data Processing and Annotation), an R package that offers comprehensive and streamlined metabolic feature extraction and annotation. | 2022-03 | 0 | 0.0 | 0.0 |  | 0 |
| Totoro | Endocrinology and metabolism Metabolomics Molecular interactions, pathways and networks Small molecules | Identifying Active Reactions During the Transient State for Metabolic Perturbations. | 2022-02 | 0 | 0.0 | 0.0 |  | 0 |
| CCDB | Endocrinology and metabolism Metabolomics Molecular interactions, pathways and networks Nutritional science Small molecules | A database for exploring inter-chemical correlations in metabolomics and exposomics datasets. | 2022-06 | 0 | 0.0 | 0.0 |  | 0 |
| Gene-SCOUT | Biobank Exome sequencing Genomics Genotype and phenotype Metabolomics | Identifying genes with similar continuous trait fingerprints from phenome-wide association analyses. | 2022-05 | 0 | 0.0 | 0.0 |  | 0 |
| BioTransformer | Carbohydrates Endocrinology and metabolism Metabolomics NMR Small molecules | BioTransformer is a freely available web server that supports accurate, rapid and comprehensive in silico metabolism prediction. | 2022-07 | 0 | 0.0 | 0.0 |  | 0 |
| DeepHisCoM | DNA polymorphism Metabolomics Metagenomics Molecular interactions, pathways and networks Transcriptomics | DeepHisCoM (Deep-learning pathway analysis using Hierarchical structured CoMponent models) which uses deep learning analysis to find complex relationships between biological factors and pathways. | -- | 0 | 0.0 | 0.0 |  | 0 |
| SeMa-Trap | Metabolomics RNA-Seq Small molecules Transcriptomics Workflows | Expression-based exploration tool for increased secondary metabolite production in bacteria. | 2022-07 | 0 | 0.0 | 0.0 |  | 0 |
| pseudoDrift | Metabolomics NMR Plant biology Proteomics experiment Small molecules | Normalizing and Correcting Variable and Complex LC-MS Metabolomic Data with the R Package pseudoDrift. | 2022-05 | 0 | 0.0 | 0.0 |  | 0 |
| ERMer | Endocrinology and metabolism Gene regulation Metabolomics Molecular interactions, pathways and networks Transcription factors and regulatory sites | ERMer: A serverless platform for navigating, analyzing, and visualizing Escherichiacoli regulatory landscape through graph database. | 2022-07 | 0 | 0.0 | 0.0 |  | 0 |
| MINE | Biochemistry Data mining Endocrinology and metabolism Metabolomics Small molecules | Enhanced biochemical coverage for peak identification in untargeted metabolomics. | 2022-07 | 0 | 0.0 | 0.0 |  | 0 |
| MACE | Biological databases Ecology Metabolomics Plant biology Proteomics experiment | An Open Access Data Repository of Mass Spectra for Chemical Ecology. | -- | 0 | 0.0 | 0.0 |  | 0 |
| ABCstats | Endocrinology and metabolism Mathematics Metabolomics Small molecules | ABCstats is an R package for metabolomics data transformation. Adaptive Box-Cox (ABC) transformation improves the data normality for statistical analysis. | 2022-06 | 0 | 0.0 | 0.0 |  | 0 |
| semiquantification | Metabolomics Proteomics experiment Small molecules Workflows | Novel Workflow for Semiquantification of Emerging Contaminants in Environmental Samples Analyzed by Gas Chromatography-Atmospheric Pressure Chemical Ionization-Quadrupole Time of Flight-Mass Spectrometry. | 2022-07 | 0 | 0.0 | 0.0 |  | 0 |
| QLattice | Biomarkers Machine learning Metabolomics Molecular interactions, pathways and networks Proteomics | Identifying interactions in omics data for clinical biomarker discovery using symbolic regression. | 2022-08 | 0 | 0.0 | 0.0 |  | 0 |
| PeakForest | Data submission, annotation and curation Metabolomics NMR Proteomics experiment Small molecules | A multi-platform digital infrastructure for interoperable metabolite spectral data and metadata management. | 2022-06 | 0 | 0.0 | 0.0 |  | 0 |
| MiMIR | Biobank Endocrinology and metabolism Metabolomics Public health and epidemiology Small molecules | R-shiny application to infer risk factors and endpoints from Nightingale Healths 1H-NMR Metabolomics data. | 2022-08 | 0 | 0.0 | 0.0 |  | 0 |
| MIAOME | Epigenomics Functional, regulatory and non-coding RNA Metabolomics Microbial ecology Small molecules | Human microbiome affect the host epigenome. | 2022-01 | 0 | 0.0 | 0.0 |  | 0 |
| mGWAS-Explorer | DNA polymorphism GWAS study Metabolomics Pathology Small molecules | Linking SNPs, Genes, Metabolites, and Diseases for Functional Insights. | 2022-06 | 0 | 0.0 | 0.0 |  | 0 |
| metapone | Endocrinology and metabolism Metabolomics Molecular interactions, pathways and networks Proteomics experiment Small molecules | A Bioconductor package for joint pathway testing for untargeted metabolomics data. | 2022-07 | 0 | 0.0 | 0.0 |  | 0 |
| massDatabase | Endocrinology and metabolism Lipids Metabolomics Molecular interactions, pathways and networks Small molecules | massdatabase is an R package that operates the online public databases and combines with other tools for streamlined compound annotation and pathway enrichment analysis. massDatabase is a flexible, simple, and powerful tool that can be installed on all platforms, allowing the users to leverage all the online public databases for biological function mining. | -- | 0 | 0.0 | 0.0 |  | 0 |
| ConSIG | Biomarkers Metabolomics Proteomics Sequence analysis Transcriptomics | ConSIG is a web page which can discovery of molecular signature from OMIC data. | 2022-07 | 0 | 0.0 | 0.0 |  | 0 |
| GATOM | Endocrinology and metabolism Lipids Metabolomics Molecular interactions, pathways and networks Small molecules | Omics-based identification of regulated metabolic modules in atom transition networks. | 2022-07 | 0 | 0.0 | 0.0 |  | 0 |
| ToxSTAR | Biotechnology Metabolomics Pathology Small molecules Toxicology | ToxSTAR is a platform to predict human toxicity caused by drugs and chemicals through integration of biotechnology (BT) and information technology (IT). | -- | 0 | 0.0 | 0.0 |  | 0 |
| REGLIV | Gene and protein families Gene regulation Metabolomics Oncology Proteomics | Molecular regulation data of diverse living systems facilitating current multiomics research. | 2022-09 | 0 | 0.0 | 0.0 |  | 0 |
| MRSCloud | Metabolomics Molecular interactions, pathways and networks NMR Small molecules | A cloud-based spectral simulation tool “MRSCloud,” which allows MRS users to simulate a vendor-specific and sequence-specific basis set online in a convenient and time-efficient manner. This tool can simulate basis sets for GE, Philips, and Siemens MR scanners, including conventional acquisitions and spectral editing schemes with PRESS and semi-LASER localization at 3 T. | 2022-11 | 0 | 0.0 | 0.0 |  | 0 |
| MobilityTransformR | Bioinformatics Metabolomics Workflows | An R package for effective mobility transformation of CE-MS data. | 2022-08 | 0 | 0.0 | 0.0 |  | 0 |
| deepGraphh | Cheminformatics Machine learning Metabolomics Microbial ecology Small molecules | DeepGraphh is an open-source web service that features a conglomerate of powerful graph-based neural networks methods, with highly configurable parameters support via the graphical user interface (GUI). | -- | 0 | 0.0 | 0.0 |  | 0 |
| binneR | Biomarkers Metabolomics NMR Proteomics experiment | A spectral binning approach for flow infusion electrospray high resolution mass spectrometry (FIE-HRMS) metabolome fingerprinting data. | 2022-08 | 0 | 0.0 | 0.0 |  | 0 |
| CPRiL | Literature and language Metabolomics Molecular biology Small molecules | CPRiL is a webserver for exploring functional compound-protein relationships that are extracted automatically from PubMed literature. | -- | 0 | 0.0 | 0.0 |  | 0 |
| NPvis | Lipids Metabolomics Proteomics Proteomics experiment Small molecules | An Interactive Visualizer of Peptidic Natural Product-MS/MS Matches. | 2022-08 | 0 | 0.0 | 0.0 |  | 0 |
| IDSL.UFA | Chemistry Metabolomics Molecular interactions, pathways and networks Proteomics experiment Workflows | IDSL.UFA Assigns High-Confidence Molecular Formula Annotations for Untargeted LC/HRMS Data Sets in Metabolomics and Exposomics. | 2022-10 | 0 | 0.0 | 0.0 |  | 0 |
| TurboPutative | Carbohydrates Lipids Metabolomics Molecular interactions, pathways and networks Small molecules | A web server for data handling and metabolite classification in untargeted metabolomics. | 2022-09 | 0 | 0.0 | 0.0 |  | 0 |
| dbRUSP | Endocrinology and metabolism Metabolomics Nutritional science Proteomics experiment Small molecules | DbRUSP is an interactive database for the analysis and interpretation of newborn screening (NBS) data and for studying the effects of covariates on blood metabolite levels. | 2022-09 | 0 | 0.0 | 0.0 |  | 0 |
| KRiShI | Literature and language Metabolomics Molecular interactions, pathways and networks Pathology Plant biology | Knowledgebase for rice sheath blight information (KRiShI) is a manually curated user-friendly knowledgebase for rice sheath blight (SB) disease that allows users to efficiently mine, visualize, search, benchmark, download, and update meaningful data and information related to SB using its easy and interactive interface. | 2022-01 | 0 | 0.0 | 0.0 |  | 0 |
| metabolomicsR | Metabolomics Proteomics Proteomics experiment Small molecules Workflows | MetabolomicsR is a streamlined R package to preprocess, analyze, and visualize metabolomic data. | -- | 0 | 0.0 | 0.0 |  | 0 |
| MIMaL | Machine learning Metabolomics Proteomics Proteomics experiment Small molecules | The Multi-Omic Integration by Machine Learning (MIMaL) method can be found as scripts and a web page. | 2022-10 | 0 | 0.0 | 0.0 |  | 0 |